6726 results
X-ray diffraction data for the Crystal structure of Transaldolase (EC 2.2.1.2) (TM0295) from Thermotoga maritima at 2.40 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.40 Å
R/Rfree: 0.20/0.25
Resolution: 2.40 Å
R/Rfree: 0.20/0.25
X-ray diffraction data for the Crystal structure of Activation (AdoMet binding) domain of Methionine synthase (TM0269) from Thermotoga maritima at 2.2 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.30 Å
R/Rfree: 0.25/0.29
Resolution: 2.30 Å
R/Rfree: 0.25/0.29
X-ray diffraction data for the Crystal structure of DNA polymerase III, beta subunit (TM0262) from Thermotoga maritima at 2.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.20/0.23
Resolution: 2.00 Å
R/Rfree: 0.20/0.23
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE RNFG SUBUNIT OF ELECTRON TRANSPORT COMPLEX (TM0246) FROM THERMOTOGA MARITIMA AT 1.65 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.65 Å
R/Rfree: 0.18/0.21
Resolution: 1.65 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of Glycyl-tRNA synthetase alpha chain (TM0216) from Thermotoga maritima at 1.95 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.95 Å
R/Rfree: 0.20/0.25
Resolution: 1.95 Å
R/Rfree: 0.20/0.25
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE ENDORIBONUCLEASE (TM0215) FROM THERMOTOGA MARITIMA MSB8 AT 2.30 A RESOLUTION

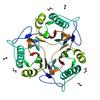
First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.30 Å
R/Rfree: 0.20/0.26
Resolution: 2.30 Å
R/Rfree: 0.20/0.26
X-ray diffraction data for the Crystal structure of Glycine cleavage system H protein (tm0212) from Thermotoga maritima at 1.65 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.65 Å
R/Rfree: 0.23/0.27
Resolution: 1.65 Å
R/Rfree: 0.23/0.27
X-ray diffraction data for the Crystal structure of a putative zn-dependent hydrolase of the metallo-beta-lactamase superfamily (tm0207) from thermotoga maritima at 2.00 A resolution

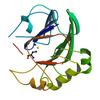
First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.18/0.22
Resolution: 2.00 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of a putative fe(iii) abc transporter (tm0189) from thermotoga maritima msb8 at 1.70 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.16/0.21
Resolution: 1.70 Å
R/Rfree: 0.16/0.21
X-ray diffraction data for the Crystal structure of Acyl carrier protein (TM0175) from Thermotoga maritima at 2.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.20/0.25
Resolution: 2.00 Å
R/Rfree: 0.20/0.25
X-ray diffraction data for the Crystal structure of Folylpolyglutamate synthase (TM0166) from Thermotoga maritima at 2.10 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.10 Å
R/Rfree: 0.18/0.24
Resolution: 2.10 Å
R/Rfree: 0.18/0.24
X-ray diffraction data for the Crystal structure of Geranyltranstransferase (EC 2.5.1.10) (tm0161) from THERMOTOGA MARITIMA at 1.90 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.17/0.20
Resolution: 1.90 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE XANTHOSINE TRIPHOSPHATE PYROPHOSPHATASE/HAM1 PROTEIN HOMOLOG (TM0159) FROM THERMOTOGA MARITIMA AT 1.78 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.78 Å
R/Rfree: 0.17/0.21
Resolution: 1.78 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of indole-3-glycerol phosphate synthase (TM0140) from Thermotoga maritima at 3.0 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 3.00 Å
R/Rfree: 0.24/0.30
Resolution: 3.00 Å
R/Rfree: 0.24/0.30
X-ray diffraction data for the Crystal structure of a putative acetamidase (tm0119) from thermotoga maritima msb8 at 2.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.50 Å
R/Rfree: 0.19/0.24
Resolution: 2.50 Å
R/Rfree: 0.19/0.24
X-ray diffraction data for the Crystal structure of Acetyl xylan esterase (TM0077) from THERMOTOGA MARITIMA at 2.12 A resolution (paraoxon inhibitor complex structure)


First author:
M. Levisson
Resolution: 2.12 Å
R/Rfree: 0.17/0.21
Resolution: 2.12 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of Acetyl xylan esterase (TM0077) from THERMOTOGA MARITIMA at 2.40 A resolution (PMSF inhibitor complex structure)


First author:
M. Levisson
Resolution: 2.40 Å
R/Rfree: 0.16/0.21
Resolution: 2.40 Å
R/Rfree: 0.16/0.21
X-ray diffraction data for the Crystal structure of Acetyl xylan esterase (TM0077) from THERMOTOGA MARITIMA at 2.50 A resolution (native apo structure)


First author:
M. Levisson
Resolution: 2.50 Å
R/Rfree: 0.17/0.21
Resolution: 2.50 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of Acetyl xylan esterase (TM0077) from Thermotoga maritima at 2.10 A resolution


First author:
M. Levisson
Resolution: 2.10 Å
R/Rfree: 0.18/0.22
Resolution: 2.10 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of 2-dehydro-3- deoxygluconokinase (EC 2.7.1.45) (tm0067) from THERMOTOGA MARITIMA at 2.05 A resolution


First author:
I.I. Mathews
Resolution: 2.05 Å
R/Rfree: 0.18/0.23
Resolution: 2.05 Å
R/Rfree: 0.18/0.23
X-ray diffraction data for the Crystal structure of 2-dehydro-3-deoxyphosphogluconate aldolase/4-hydroxy-2-oxoglutarate aldolase (TM0066) from Thermotoga maritima at 2.30 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.30 Å
R/Rfree: 0.18/0.22
Resolution: 2.30 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of uronate isomerase (TM0064) from Thermotoga maritima at 2.85 A resolution


First author:
R. Schwarzenbacher
Resolution: 2.85 Å
R/Rfree: 0.23/0.27
Resolution: 2.85 Å
R/Rfree: 0.23/0.27
X-ray diffraction data for the Crystal structure of a putative PII-like signaling protein (TM0021) from Thermotoga maritima at 2.50 A resolution


First author:
R. Schwarzenbacher
Resolution: 2.50 Å
R/Rfree: 0.18/0.23
Resolution: 2.50 Å
R/Rfree: 0.18/0.23
X-ray diffraction data for the Crystal structure of pyruvate oxidoreductase subunit PORC (EC 1.2.7.1) (TM0015) from Thermotoga maritima at 2.12 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.12 Å
R/Rfree: 0.22/0.25
Resolution: 2.12 Å
R/Rfree: 0.22/0.25

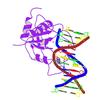
First author:
L. Halabelian
Resolution: 2.60 Å
R/Rfree: 0.21/0.25
Resolution: 2.60 Å
R/Rfree: 0.21/0.25


First author:
C. Xu
Resolution: 1.85 Å
R/Rfree: 0.20/0.24
Resolution: 1.85 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the tudor in complex with ligand


First author:
H. Zhang
Resolution: 1.93 Å
R/Rfree: 0.21/0.23
Resolution: 1.93 Å
R/Rfree: 0.21/0.23
X-ray diffraction data for the Crystal Structure of the extended TUDOR domain from TDRD2

First author:
H. Zhang
Resolution: 1.95 Å
R/Rfree: 0.19/0.22
Resolution: 1.95 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of human TBC1 domain family member 7


First author:
X. Guan
Resolution: 1.90 Å
R/Rfree: 0.21/0.26
Resolution: 1.90 Å
R/Rfree: 0.21/0.26
X-ray diffraction data for the RabGAP domain of human TBC1D22B


First author:
Y. Tong
Resolution: 2.30 Å
R/Rfree: 0.20/0.24
Resolution: 2.30 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the Trypanosoma brucei bromodomain BDF5 (Tb427tmp.01.5000)


First author:
Y.H. Lin
Resolution: 2.10 Å
R/Rfree: 0.19/0.22
Resolution: 2.10 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of a DUF1312 family protein (EF3258) from Enterococcus faecalis V583 at 1.73 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.73 Å
R/Rfree: 0.17/0.20
Resolution: 1.73 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Thebaine 6-O-demethylase (T6ODM) from Papaver somniferum in complex with 2-oxoglutarate


First author:
A. Kluza
Resolution: 1.85 Å
R/Rfree: 0.16/0.21
Resolution: 1.85 Å
R/Rfree: 0.16/0.21
X-ray diffraction data for the Thebaine 6-O-demethylase (T6ODM) from Papaver somniferum in complex with succinate


First author:
A. Kluza
Resolution: 1.97 Å
R/Rfree: 0.16/0.21
Resolution: 1.97 Å
R/Rfree: 0.16/0.21
X-ray diffraction data for the Crystal Structure of the SET Domain of Human SUV420H1 In Complex With Inhibitor


First author:
L. Halabelian
Resolution: 2.30 Å
R/Rfree: 0.23/0.27
Resolution: 2.30 Å
R/Rfree: 0.23/0.27
X-ray diffraction data for the Crystal structure of dihydrofolate reductase from Staphylococcus aureus MW2 bound to NADP and p218


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.90 Å
R/Rfree: 0.18/0.22
Resolution: 1.90 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal Structure of Streptomyces sp. SF2575 glycosyltransferase SsfS6, complexed with thymidine diphosphate


First author:
F. Wang
Resolution: 2.40 Å
R/Rfree: 0.24/0.27
Resolution: 2.40 Å
R/Rfree: 0.24/0.27
X-ray diffraction data for the Crystal Structure of SsfS6, Streptomyces sp. SF2575 glycosyltransferase


First author:
F. Wang
Resolution: 2.40 Å
R/Rfree: 0.22/0.26
Resolution: 2.40 Å
R/Rfree: 0.22/0.26
X-ray diffraction data for the Crystal structure of a N-acetylmuramoyl-L-alanine amidase (BACUNI_02947) from Bacteroides uniformis ATCC 8492 at 1.07 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.07 Å
R/Rfree: 0.11/0.14
Resolution: 1.07 Å
R/Rfree: 0.11/0.14
X-ray diffraction data for the Crystal structure of a N-acetylmuramoyl-L-alanine amidase (BACUNI_02947) from Bacteroides uniformis ATCC 8492 at 1.30 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.30 Å
R/Rfree: 0.12/0.15
Resolution: 1.30 Å
R/Rfree: 0.12/0.15
X-ray diffraction data for the Crystal structure of a N-acetylmuramoyl-L-alanine amidase (BACUNI_02947) from Bacteroides uniformis ATCC 8492 at 1.15 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.15 Å
R/Rfree: 0.13/0.16
Resolution: 1.15 Å
R/Rfree: 0.13/0.16
X-ray diffraction data for the Crystal structure of a putative GDSL-like lipase (BACOVA_00914) from Bacteroides ovatus ATCC 8483 at 1.40 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.40 Å
R/Rfree: 0.13/0.16
Resolution: 1.40 Å
R/Rfree: 0.13/0.16
X-ray diffraction data for the Crystal structure of a putative oxidoreductase (PARMER_00841) from Parabacteroides merdae ATCC 43184 at 2.23 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.23 Å
R/Rfree: 0.19/0.22
Resolution: 2.23 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of a putative carbohydrate binding protein (BACOVA_03559) from Bacteroides ovatus ATCC 8483 at 1.50 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.50 Å
R/Rfree: 0.16/0.18
Resolution: 1.50 Å
R/Rfree: 0.16/0.18
X-ray diffraction data for the Crystal structure of a putative adhesin (BACEGG_01763) from Bacteroides eggerthii DSM 20697 at 2.20 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.20 Å
R/Rfree: 0.21/0.24
Resolution: 2.20 Å
R/Rfree: 0.21/0.24
X-ray diffraction data for the Crystal structure of a thioredoxin-like protein (BACCAC_02376) from Bacteroides caccae ATCC 43185 at 1.90 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.18/0.21
Resolution: 1.90 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of a susD homolog (BT1281) from Bacteroides thetaiotaomicron VPI-5482 at 1.90 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.14/0.18
Resolution: 1.90 Å
R/Rfree: 0.14/0.18
X-ray diffraction data for the Crystal structure of a susd homolog (BACOVA_04803) from Bacteroides ovatus ATCC 8483 at 2.25 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.25 Å
R/Rfree: 0.15/0.17
Resolution: 2.25 Å
R/Rfree: 0.15/0.17
X-ray diffraction data for the Crystal structure of a SusD homolog (BACUNI_02643) from Bacteroides uniformis ATCC 8492 at 2.00 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.17/0.20
Resolution: 2.00 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of a putative epoxide hydrolase (KPN_01808) from Klebsiella pneumoniae subsp. pneumoniae MGH 78578 at 1.60 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.60 Å
R/Rfree: 0.15/0.17
Resolution: 1.60 Å
R/Rfree: 0.15/0.17
X-ray diffraction data for the Crystal structure of a Putative efflux transporter (BACEGG_01895) from Bacteroides eggerthii DSM 20697 at 2.06 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.06 Å
R/Rfree: 0.21/0.23
Resolution: 2.06 Å
R/Rfree: 0.21/0.23
X-ray diffraction data for the Crystal structure of a similar to lipoprotein, NLP/P60 family (SAV0400) from Staphylococcus aureus subsp. aureus Mu50 at 2.23 A resolution


First author:
Q. Xu
Resolution: 2.23 Å
R/Rfree: 0.20/0.24
Resolution: 2.23 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the Crystal structure of a putative dipeptidyl-peptidase VI (BACOVA_00612) from Bacteroides ovatus at 1.72 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.72 Å
R/Rfree: 0.14/0.17
Resolution: 1.72 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of a clostripain (PARMER_00083) from Parabacteroides merdae ATCC 43184 at 1.70 A resolution


First author:
K. McLuskey
Resolution: 1.70 Å
R/Rfree: 0.14/0.18
Resolution: 1.70 Å
R/Rfree: 0.14/0.18
X-ray diffraction data for the Crystal structure of a putative lipase (lip1) from Acinetobacter baumannii AYE at 1.70 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.16/0.19
Resolution: 1.70 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of a putative acylhydrolase (BACUNI_03406) from Bacteroides uniformis ATCC 8492 at 1.37 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.37 Å
R/Rfree: 0.13/0.17
Resolution: 1.37 Å
R/Rfree: 0.13/0.17
X-ray diffraction data for the Crystal structure of a putative alpha-galactosidase/melibiase (BF4189) from Bacteroides fragilis NCTC 9343 at 2.00 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.17/0.20
Resolution: 2.00 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of a putative alpha-galactosidase (BF1418) from Bacteroides fragilis NCTC 9343 at 1.57 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.57 Å
R/Rfree: 0.15/0.17
Resolution: 1.57 Å
R/Rfree: 0.15/0.17
X-ray diffraction data for the Crystal structure of a leucine-rich repeat protein (EUBVEN_01088) from Eubacterium ventriosum ATCC 27560 at 2.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.50 Å
R/Rfree: 0.18/0.21
Resolution: 2.50 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of an ankyrin repeat domain (ABAYE2397) from Acinetobacter baumannii AYE at 1.00 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.00 Å
R/Rfree: 0.14/0.17
Resolution: 1.00 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of a DUF4785 family protein (lpg0956) from Legionella pneumophila subsp. pneumophila str. Philadelphia 1 at 2.00 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.19/0.22
Resolution: 2.00 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of a putative nutrient binding protein (lpg2210) from Legionella pneumophila subsp. pneumophila str. Philadelphia 1 at 2.33 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.33 Å
R/Rfree: 0.17/0.22
Resolution: 2.33 Å
R/Rfree: 0.17/0.22
X-ray diffraction data for the Crystal structure of an uncharacterized protein (EUBREC_2869) from Eubacterium rectale ATCC 33656 at 1.45 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.45 Å
R/Rfree: 0.16/0.19
Resolution: 1.45 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of a putative secreted protein (EUBREC_3654) from Eubacterium rectale at 2.43 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.43 Å
R/Rfree: 0.19/0.22
Resolution: 2.43 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of an uncharacterized beta-sandwich protein (CD630_09570) from Clostridium difficile 630 at 1.90 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.17/0.19
Resolution: 1.90 Å
R/Rfree: 0.17/0.19
X-ray diffraction data for the Crystal structure of a putative lipoprotein (CD1622) from Clostridium difficile 630 at 1.85 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.85 Å
R/Rfree: 0.18/0.21
Resolution: 1.85 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of a DUF2790 family protein (PA3229) from Pseudomonas aeruginosa PAO1 at 1.26 A resolution

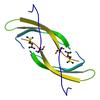
First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.26 Å
R/Rfree: 0.14/0.17
Resolution: 1.26 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of a DUF4398 family protein (PA2901) from Pseudomonas aeruginosa PAO1 at 2.47 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.47 Å
R/Rfree: 0.20/0.24
Resolution: 2.47 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the Crystal structure of a DUF3299 family protein (PA4066) from Pseudomonas aeruginosa PAO1 at 2.45 A resolution

First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.45 Å
R/Rfree: 0.17/0.21
Resolution: 2.45 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of an IcmL-like type IV secretion system protein (lpg0120) from Legionella pneumophila subsp. pneumophila str. Philadelphia 1 at 2.65 A resolution

First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.65 Å
R/Rfree: 0.19/0.22
Resolution: 2.65 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of a putative secreted protein (lpg1979) from Legionella pneumophila subsp. pneumophila str. Philadelphia 1 at 2.65 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.65 Å
R/Rfree: 0.16/0.20
Resolution: 2.65 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the Crystal structure of a DUF3298 family protein (PA4972) from Pseudomonas aeruginosa PAO1 at 2.15 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.15 Å
R/Rfree: 0.17/0.21
Resolution: 2.15 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of a putative hydrolase (lpg1103) from Legionella pneumophila subsp. pneumophila str. Philadelphia 1 at 1.15 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.15 Å
R/Rfree: 0.14/0.16
Resolution: 1.15 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the Crystal structure of a binding protein component of ABC phosphonate transporter (PA3383) from Pseudomonas aeruginosa at 1.97 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.97 Å
R/Rfree: 0.16/0.19
Resolution: 1.97 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of a putative carbohydrate binding protein (PA1324) from Pseudomonas aeruginosa at 2.61 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.61 Å
R/Rfree: 0.21/0.23
Resolution: 2.61 Å
R/Rfree: 0.21/0.23
X-ray diffraction data for the Crystal structure of a DUF4426 family protein (PA0388) from Pseudomonas aeruginosa PAO1 at 2.09 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.09 Å
R/Rfree: 0.21/0.23
Resolution: 2.09 Å
R/Rfree: 0.21/0.23
X-ray diffraction data for the Crystal structure of a fimbrial biogenesis protein PilY2 (PilY2_PA4555) from Pseudomonas aeruginosa PAO1 at 2.10 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.10 Å
R/Rfree: 0.19/0.22
Resolution: 2.10 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of an inhibitor of vertebrate lysozyme (PA3902) from Pseudomonas aeruginosa PAO1 at 1.25 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.25 Å
R/Rfree: 0.12/0.16
Resolution: 1.25 Å
R/Rfree: 0.12/0.16
X-ray diffraction data for the Crystal structure of a protein with unknown function from DUF2059 family (PA0856) from PSEUDOMONAS AERUGINOSA at 2.72 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.72 Å
R/Rfree: 0.21/0.24
Resolution: 2.72 Å
R/Rfree: 0.21/0.24
X-ray diffraction data for the Crystal structure of a DUF4466 family protein (PARMER_03218) from Parabacteroides merdae ATCC 43184 at 2.00 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.17/0.22
Resolution: 2.00 Å
R/Rfree: 0.17/0.22
X-ray diffraction data for the Crystal structure of an ABC-type phosphate transport system, periplasmic component (LVIS_0633) from Lactobacillus brevis ATCC 367 at 1.55 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.55 Å
R/Rfree: 0.16/0.20
Resolution: 1.55 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the Crystal structure of a DUF4822 family protein (EF0375) from Enterococcus faecalis V583 at 1.85 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.85 Å
R/Rfree: 0.15/0.18
Resolution: 1.85 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of a Lipocalin-like protein (EF0376) from Enterococcus faecalis V583 at 1.60 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.60 Å
R/Rfree: 0.14/0.17
Resolution: 1.60 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of an uncharacterized protein (PARMER_03598) from Parabacteroides merdae ATCC 43184 at 1.50 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.50 Å
R/Rfree: 0.17/0.19
Resolution: 1.50 Å
R/Rfree: 0.17/0.19
X-ray diffraction data for the Crystal structure of a modulator protein MzrA (KPN_03524) from Klebsiella pneumoniae subsp. pneumoniae MGH 78578 at 2.45 A resolution

First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.45 Å
R/Rfree: 0.20/0.25
Resolution: 2.45 Å
R/Rfree: 0.20/0.25
X-ray diffraction data for the Crystal structure of an alpha-lytic protease prodomain-like protein (DESPIG_01740) from Desulfovibrio piger ATCC 29098 at 1.80 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.80 Å
R/Rfree: 0.20/0.22
Resolution: 1.80 Å
R/Rfree: 0.20/0.22
X-ray diffraction data for the Crystal structure of a DUF487 family protein (DESPIG_00776) from Desulfovibrio piger ATCC 29098 at 2.49 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.40 Å
R/Rfree: 0.18/0.23
Resolution: 2.40 Å
R/Rfree: 0.18/0.23
X-ray diffraction data for the Crystal structure of an Acid stress chaperone HdeB (KPN_03484) from Klebsiella pneumoniae subsp. pneumoniae MGH 78578 at 1.70 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.20/0.23
Resolution: 1.70 Å
R/Rfree: 0.20/0.23
X-ray diffraction data for the Crystal structure of a DUF1329 family protein (DESPIG_00262) from Desulfovibrio piger ATCC 29098 at 1.75 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.75 Å
R/Rfree: 0.21/0.25
Resolution: 1.75 Å
R/Rfree: 0.21/0.25
X-ray diffraction data for the Crystal structure of a putative lipoprotein from the DUF903 family (KPN_03160) from Klebsiella pneumoniae subsp. pneumoniae MGH 78578 at 2.30 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.30 Å
R/Rfree: 0.18/0.22
Resolution: 2.30 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of a thioesterase-like protein (CLOSPO_01618) from Clostridium sporogenes ATCC 15579 at 1.70 A resolution

First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.18/0.20
Resolution: 1.70 Å
R/Rfree: 0.18/0.20
X-ray diffraction data for the Crystal structure of a DUF4909 family protein (SAV1798) from Staphylococcus aureus subsp. aureus Mu50 at 1.92 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.92 Å
R/Rfree: 0.20/0.24
Resolution: 1.92 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the Crystal structure of a protein with alpha-lytic protease prodomain-like fold (BDI_0842) from Parabacteroides distasonis ATCC 8503 at 1.40 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.40 Å
R/Rfree: 0.13/0.17
Resolution: 1.40 Å
R/Rfree: 0.13/0.17
X-ray diffraction data for the Crystal structure of a TPR-domain containing protein (BDI_1685) from Parabacteroides distasonis ATCC 8503 at 2.26 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.26 Å
R/Rfree: 0.18/0.21
Resolution: 2.26 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of a putative esterase (BDI_1566) from Parabacteroides distasonis ATCC 8503 at 1.60 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.60 Å
R/Rfree: 0.16/0.19
Resolution: 1.60 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of a putative cell adhesion protein (CLOSPO_03726) from Clostridium sporogenes ATCC 15579 at 1.95 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.95 Å
R/Rfree: 0.17/0.21
Resolution: 1.95 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of a conserved domain protein (SP_1775) from Streptococcus pneumoniae TIGR4 at 1.91 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.91 Å
R/Rfree: 0.18/0.23
Resolution: 1.91 Å
R/Rfree: 0.18/0.23
X-ray diffraction data for the Crystal structure of a YbbR-like protein (SP_1560) from Streptococcus pneumoniae TIGR4 at 2.74 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.74 Å
R/Rfree: 0.23/0.29
Resolution: 2.74 Å
R/Rfree: 0.23/0.29
X-ray diffraction data for the Crystal structure of a DUF4467 family protein (SAV0303) from Staphylococcus aureus subsp. aureus Mu50 at 1.35 A resolution

First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.20 Å
R/Rfree: 0.14/0.17
Resolution: 1.20 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of a cysteine-rich secretory protein (SAV1118) from Staphylococcus aureus subsp. aureus Mu50 at 1.90 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.19/0.22
Resolution: 1.90 Å
R/Rfree: 0.19/0.22