6617 results
X-ray diffraction data for the Crystal structure of trypsin at 100 Kelvin with benzamidine

X-ray diffraction data for the Crystal structure of trypsin at 125 Kelvin with benzamidine
X-ray diffraction data for the Crystal structure of trypsin at 125 Kelvin with benzamidine (duplicate)

X-ray diffraction data for the Crystal structure of trypsin at 125 Kelvin with benzamidine (triplicate)

X-ray diffraction data for the Crystal structure of trypsin at 100 Kelvin with benzamidine (Duplicate)

X-ray diffraction data for the Crystal structure of trypsin at 150 Kelvin with benzamidine (triplicate)

X-ray diffraction data for the Crystal structure of trypsin at 150 Kelvin with benzamidine (duplicate)

X-ray diffraction data for the Crystal structure of trypsin at 150 Kelvin with benzamidine
X-ray diffraction data for the Crystal structure of trypsin at 175 Kelvin with benzamidine
X-ray diffraction data for the Crystal structure of trypsin at 200 Kelvin with benzamidine (duplicate)

X-ray diffraction data for the Crystal structure of trypsin at 175 Kelvin with benzamidine (duplicate)

X-ray diffraction data for the Crystal structure of trypsin at 200 Kelvin with benzamidine

X-ray diffraction data for the Crystal structure of trypsin at 200 Kelvin with benzamidine (triplicate)

X-ray diffraction data for the Crystal structure of trypsin at 175 Kelvin with benzamidine (triplicate)

X-ray diffraction data for the Crystal structure of trypsin at 225 Kelvin with benzamidine (Triplicate)

X-ray diffraction data for the Crystal structure of trypsin at 225 Kelvin with benzamidine

X-ray diffraction data for the Crystal structure of trypsin at 250 Kelvin with benzamidine

X-ray diffraction data for the Crystal structure of trypsin at 250 Kelvin with benzamidine (duplicate)

X-ray diffraction data for the Crystal structure of trypsin at 225 Kelvin with benzamidine (duplicate)

X-ray diffraction data for the Crystal structure of trypsin at 275 Kelvin with benzamidine (duplicate)
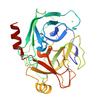
X-ray diffraction data for the Crystal structure of trypsin at 275 Kelvin with benzamidine (triplicate)

X-ray diffraction data for the Crystal structure of trypsin at 275 Kelvin with benzamidine

X-ray diffraction data for the Crystal structure of trypsin at 250 Kelvin with benzamidine (triplicate)

X-ray diffraction data for the Co-structure of the Fab of the anti-TIGIT Vibostolimab antibody with its antigen


X-ray diffraction data for the Crystal Structure of GTP cyclohydrolase 1 (FolE) from Mycobacterium tuberculosis


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: P9WN57
Resolution: 2.74 Å
R/Rfree: 0.21/0.26
Uniprot: P9WN57
Resolution: 2.74 Å
R/Rfree: 0.21/0.26
X-ray diffraction data for the Crystal structure of trypsin at 300 Kelvin with benzamidine

X-ray diffraction data for the Crystal structure of trypsin at 300 Kelvin with benzamidine (duplicate)

X-ray diffraction data for the Crystal structure of the SpyTag003-SpyCatcher003 complex


X-ray diffraction data for the Crystal structure of the allosteric MKP5 mutant Y435W


X-ray diffraction data for the Crystal structure of an MKP5 mutant, Y435W, in complex with an allosteric inhibitor


X-ray diffraction data for the Crystal structure of HP1alpha chromoshadow domain in complex with KAP1 peptide


X-ray diffraction data for the Crystal structure of HP1alpha chromoshadow domain


X-ray diffraction data for the Helicobacter pylori cysteine rich protein B (hcpB)

X-ray diffraction data for the Crystallizing extracellular protein reveals paths for silver mineralization and recovery
First author:
Fiaz Ahmad
X-ray diffraction data for the A Stable Heterotrimeric Foldon with Minimum Mutation


X-ray diffraction data for the Crystal structure of HIV-1 Reverse Transcriptase RNase H domain complexed with a galloyl inhibitor

X-ray diffraction data for the Crystal structure of trypsin at 100 Kelvin with benzamidine (triplicate)

X-ray diffraction data for the Crystal structure of ENL YEATS in complex with histone H3 methacrylated at K18

X-ray diffraction data for the Crystal structure of the H3 hemagglutinin COBRA TJ2


X-ray diffraction data for the Crystal structure of the H3 hemagglutinin COBRA J4


X-ray diffraction data for the Crystal structure of nucleoside-diphosphate kinase from Cryptosporidium parvum in complex with citrate


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: Q5CR64
Resolution: 1.45 Å
R/Rfree: 0.13/0.17
Uniprot: Q5CR64
Resolution: 1.45 Å
R/Rfree: 0.13/0.17
X-ray diffraction data for the Anti-HIV-1 chimeric miniprotein mimicking the N-terminal half of gp41 NHR with an extended region targeting the MPER


X-ray diffraction data for the The ubiquitin-associated domain of human thirty-eight negative kinase 1, fused to the 3TEL crystallization chaperone via a 2-glycine linker


X-ray diffraction data for the Crystal structure of 2,3,4,5-tetrahydropyridine-2,6-dicarboxylate N-succinyltransferase (DapD) from Bordetella pertussis (succinyl-CoA bound)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: P0A4U8
Resolution: 1.45 Å
R/Rfree: 0.16/0.18
Uniprot: P0A4U8
Resolution: 1.45 Å
R/Rfree: 0.16/0.18
X-ray diffraction data for the Crystal structure of UTP--glucose-1-phosphate uridylyltransferase from Bordetella pertussis


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: Q7VTV0
Resolution: 1.93 Å
R/Rfree: 0.18/0.21
Uniprot: Q7VTV0
Resolution: 1.93 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of nucleoside-diphosphate kinase Cryptosporidium parvum


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: Q5CR64
Resolution: 1.31 Å
R/Rfree: 0.15/0.19
Uniprot: Q5CR64
Resolution: 1.31 Å
R/Rfree: 0.15/0.19
X-ray diffraction data for the Crystal structure of nucleoside-diphosphate kinase from Cryptosporidium parvum in complex with thymidine-5'-phosphate


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: Q5CR64
Resolution: 1.51 Å
R/Rfree: 0.14/0.16
Uniprot: Q5CR64
Resolution: 1.51 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the Crystal structure of nucleoside-diphosphate kinase from Cryptosporidium parvum in complex with cytidine-5'-diphosphate and cytidine-5'-triphosphate


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: Q5CR64
Resolution: 1.61 Å
R/Rfree: 0.14/0.17
Uniprot: Q5CR64
Resolution: 1.61 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of nucleoside-diphosphate kinase from Cryptosporidium parvum in complex with with guanosine-5'-diphosphate and AMP-PNP


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: Q5CR64
Resolution: 1.63 Å
R/Rfree: 0.15/0.17
Uniprot: Q5CR64
Resolution: 1.63 Å
R/Rfree: 0.15/0.17
X-ray diffraction data for the Crystal structure of nucleoside-diphosphate kinase from Cryptosporidium parvum in complex with with ADP and CTP


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: Q5CR64
Resolution: 1.60 Å
R/Rfree: 0.15/0.18
Uniprot: Q5CR64
Resolution: 1.60 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of nucleoside-diphosphate kinase from Cryptosporidium parvum containing phosphorylated active site histidine


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: Q5CR64
Resolution: 1.58 Å
R/Rfree: 0.14/0.17
Uniprot: Q5CR64
Resolution: 1.58 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of nucleoside-diphosphate kinase from Cryptosporidium parvum in complex with with ADP


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: Q5CR64
Resolution: 1.62 Å
R/Rfree: 0.15/0.18
Uniprot: Q5CR64
Resolution: 1.62 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of Dsk2 Sti1 domain bound to a transmembrane domain


X-ray diffraction data for the Crystal Structure of Tryptophanyl-tRNA synthetase from Klebsiella aerogenes (tryptophan bound)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A0A0H3FKK2
Resolution: 2.15 Å
R/Rfree: 0.18/0.22
Uniprot: A0A0H3FKK2
Resolution: 2.15 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of 2,3,4,5-tetrahydropyridine-2,6-dicarboxylate N-succinyltransferase (DapD) from Bordetella pertussis


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: P0A4U8
Resolution: 1.59 Å
R/Rfree: 0.16/0.18
Uniprot: P0A4U8
Resolution: 1.59 Å
R/Rfree: 0.16/0.18
X-ray diffraction data for the Crystal structure of a selenomethionine-labeled hydropyrene synthase (M75L variant) in its closed conformation


X-ray diffraction data for the Crystal Structure of Tryptophanyl-tRNA synthetase from Klebsiella aerogenes


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A0A0H3FKK2
Resolution: 2.10 Å
R/Rfree: 0.20/0.25
Uniprot: A0A0H3FKK2
Resolution: 2.10 Å
R/Rfree: 0.20/0.25
X-ray diffraction data for the Crystal structure of Phosphoglycerate mutase from Trichomonas vaginalis (sulfate bound)

First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A2DUN8
Resolution: 2.05 Å
R/Rfree: 0.21/0.24
Uniprot: A2DUN8
Resolution: 2.05 Å
R/Rfree: 0.21/0.24
X-ray diffraction data for the Crystal structure of a hydropyrene synthase (M75L variant) in its closed conformation


X-ray diffraction data for the Crystal structure of Thymidylate kinase (Tmk) from Klebsiella aerogenes.


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A0A0J2I384
Resolution: 2.85 Å
R/Rfree: 0.24/0.27
Uniprot: A0A0J2I384
Resolution: 2.85 Å
R/Rfree: 0.24/0.27
X-ray diffraction data for the Crystal structure of the Plastid Redox Insensitive 2 from Arabidopsis thaliana


X-ray diffraction data for the Crystal structure of phosphoglycerate mutase from Trichomonas vaginalis


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A2DUN8
Resolution: 1.95 Å
R/Rfree: 0.16/0.19
Uniprot: A2DUN8
Resolution: 1.95 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of bifunctional farnesyl/geranylgeranyl pyrophosphate synthase (FPPS/GGPPS) from Plasmodium falciparum in complex with MMV019313


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: Q8II79
Resolution: 3.25 Å
R/Rfree: 0.25/0.29
Uniprot: Q8II79
Resolution: 3.25 Å
R/Rfree: 0.25/0.29
X-ray diffraction data for the Crystal structure of Ami1 from M. tuberculosis in complex with a tetrazole compound


X-ray diffraction data for the Crystal structure of a mutant Staphylococcus equorum manganese superoxide dismutase T89C and N187C


X-ray diffraction data for the Crystal structure of AF9 YEATS domain in complex with acetylated at K1007 MOZ


First author:
K. Selvam
Resolution: 2.30 Å
R/Rfree: 0.25/0.27
Resolution: 2.30 Å
R/Rfree: 0.25/0.27
X-ray diffraction data for the Crystal structure of AF9 YEATS domain in complex with dicrotonylated at K1007 and K1014 MOZ


First author:
K. Selvam
Resolution: 2.10 Å
R/Rfree: 0.20/0.25
Resolution: 2.10 Å
R/Rfree: 0.20/0.25
X-ray diffraction data for the Crystal structure of Purine nucleoside phosphorylase from Trichomonas vaginalis (inosine bound)

First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Resolution: 1.20 Å
R/Rfree: 0.13/0.16
Resolution: 1.20 Å
R/Rfree: 0.13/0.16
X-ray diffraction data for the Crystal structure of a bacterial clusterless MutYX bound to an Abasic site analog (THF) opposite d(8-oxo-G)


First author:
C.H. Trasvina-Arenas
Resolution: 1.55 Å
R/Rfree: 0.18/0.20
Resolution: 1.55 Å
R/Rfree: 0.18/0.20
X-ray diffraction data for the Heligmosomoides polygyrus TGF-beta Mimic 6 Domain 3 (TGM6-D3) Bound to Human TGF-beta Type II Receptor Extracellular Domain


First author:
S.E. White
Resolution: 1.40 Å
R/Rfree: 0.16/0.18
Resolution: 1.40 Å
R/Rfree: 0.16/0.18
X-ray diffraction data for the Crystal structure of HIV-1 capsid N-terminal domain in the presence of Lenacapavir


X-ray diffraction data for the Crystal structure of Purine nucleoside phosphorylase from Trichomonas vaginalis (adenosine and phosphate bound)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A2EU62
Resolution: 1.24 Å
R/Rfree: 0.13/0.15
Uniprot: A2EU62
Resolution: 1.24 Å
R/Rfree: 0.13/0.15
X-ray diffraction data for the Crystal structure of human leukocyte antigen A*0101 in complex with the Fab of alloreactive antibody E07


X-ray diffraction data for the Crystal structure of AtNATA1 bound to Ornithine and CoA


X-ray diffraction data for the Human astrovirus 2 spike bound to neutralizing scFv 4B6

First author:
S. Lanning
Resolution: 2.67 Å
R/Rfree: 0.21/0.26
Resolution: 2.67 Å
R/Rfree: 0.21/0.26
X-ray diffraction data for the Crystal structure of Purine nucleoside phosphorylase from Trichomonas vaginalis (C2 form)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A2EU62
Resolution: 1.80 Å
R/Rfree: 0.16/0.20
Uniprot: A2EU62
Resolution: 1.80 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the Crystal structure of Purine nucleoside phosphorylase from Trichomonas vaginalis (P21 form)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A2EU62
Resolution: 1.59 Å
R/Rfree: 0.15/0.18
Uniprot: A2EU62
Resolution: 1.59 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of Purine nucleoside phosphorylase from Trichomonas vaginalis (Hepes bound)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A2EU62
Resolution: 1.49 Å
R/Rfree: 0.15/0.17
Uniprot: A2EU62
Resolution: 1.49 Å
R/Rfree: 0.15/0.17
X-ray diffraction data for the Tetramer Formation of the BCL11A ZF0 Domain


X-ray diffraction data for the Crystal Structure Pyrophosphate-fructose 6-phosphate 1-phosphotransferase 1 (Pfk1) from Trichomonas vaginalis (ATP/Pyrophosphate complex)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: O61068
Resolution: 2.16 Å
R/Rfree: 0.20/0.23
Uniprot: O61068
Resolution: 2.16 Å
R/Rfree: 0.20/0.23
X-ray diffraction data for the Crystal Structure of Pyrophosphate-fructose 6-phosphate 1-phosphotransferase 1 (Pfk1) from Trichomonas vaginalis (AMP/fructose-1,6-biphosphate complex)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: O61068
Resolution: 2.83 Å
R/Rfree: 0.20/0.24
Uniprot: O61068
Resolution: 2.83 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the Crystal Structure of Pyrophosphate-fructose 6-phosphate 1-phosphotransferase 1 (Pfk1) from Trichomonas vaginalis (AMP/alpha-D-Glucose-6-phosphate complex)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: O61068
Resolution: 2.80 Å
R/Rfree: 0.20/0.24
Uniprot: O61068
Resolution: 2.80 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the Crystal Structure Pyrophosphate-fructose 6-phosphate 1-phosphotransferase 1 (Pfk1) from Trichomonas vaginalis (AMP/beta-D-Glucose-6-phosphate complex)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: O61068
Resolution: 2.62 Å
R/Rfree: 0.19/0.23
Uniprot: O61068
Resolution: 2.62 Å
R/Rfree: 0.19/0.23
X-ray diffraction data for the Crystal Structure of macrophage migration inhibitory factor from Plasmodium knowlesi


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A0A384LDV6
Resolution: 2.75 Å
R/Rfree: 0.20/0.23
Uniprot: A0A384LDV6
Resolution: 2.75 Å
R/Rfree: 0.20/0.23
X-ray diffraction data for the Crystal Structure of Pyrophosphate-fructose 6-phosphate 1-phosphotransferase 1 (Pfk1) from Trichomonas vaginalis (ADP/5-O-phosphono-alpha-D-ribofuranose complex)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: O61068
Resolution: 2.65 Å
R/Rfree: 0.21/0.26
Uniprot: O61068
Resolution: 2.65 Å
R/Rfree: 0.21/0.26
X-ray diffraction data for the Crystal structure of Purine nucleoside phosphorylase from Trichomonas vaginalis (phosphate bound)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A2EU62
Resolution: 1.27 Å
R/Rfree: 0.14/0.16
Uniprot: A2EU62
Resolution: 1.27 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the Crystal structure of Purine nucleoside phosphorylase from Trichomonas vaginalis (glycerol bound)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A2EU62
Resolution: 1.64 Å
R/Rfree: 0.15/0.18
Uniprot: A2EU62
Resolution: 1.64 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of Purine nucleoside phosphorylase from Trichomonas vaginalis (mannose bound)

First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A2EU62
Resolution: 1.18 Å
R/Rfree: 0.13/0.15
Uniprot: A2EU62
Resolution: 1.18 Å
R/Rfree: 0.13/0.15
X-ray diffraction data for the Crystal structure of Purine nucleoside phosphorylase from Trichomonas vaginalis (citrate bound)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A2EU62
Resolution: 1.89 Å
R/Rfree: 0.16/0.19
Uniprot: A2EU62
Resolution: 1.89 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of Apo Purine nucleoside phosphorylase from Trichomonas vaginalis (H32 form)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A2EU62
Resolution: 1.20 Å
R/Rfree: 0.11/0.13
Uniprot: A2EU62
Resolution: 1.20 Å
R/Rfree: 0.11/0.13
X-ray diffraction data for the Crystal structure of Purine nucleoside phosphorylase from Trichomonas vaginalis (phosphate/adenine bound)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A2EU62
Resolution: 1.25 Å
R/Rfree: 0.12/0.15
Uniprot: A2EU62
Resolution: 1.25 Å
R/Rfree: 0.12/0.15
X-ray diffraction data for the Crystal structure of Purine nucleoside phosphorylase from Trichomonas vaginalis (Adenosine and glycine complex)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A2EU62
Resolution: 1.35 Å
R/Rfree: 0.14/0.17
Uniprot: A2EU62
Resolution: 1.35 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal Structure of Bifunctional protein GlmU from Klebsiella aerogenes


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A0A0H3FLQ9
Resolution: 3.05 Å
R/Rfree: 0.18/0.21
Uniprot: A0A0H3FLQ9
Resolution: 3.05 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the STEP (PTPN5) with active-site disulfide bond and covalent ligand bound to distal C505 and C518


X-ray diffraction data for the STEP (PTPN5) with active-site disulfide bond and allosteric-site loop shift

X-ray diffraction data for the Crystal structure of Arabidopsis N-amino acetyltransferase NATA1 bound to CoA and HEPES

X-ray diffraction data for the Crystal structure of AtNATA1 bound to Acetyl CoA


X-ray diffraction data for the Crystal structure of HLA-A*03:01 in complex with a mutant PIK3CA peptide


X-ray diffraction data for the 15-Lipoxygenase-2 V427L


X-ray diffraction data for the STEP (PTPN5) at high resolution with citrate bound in active site