402 results
X-ray diffraction data for the 2.15 Angstrom Crystal Structure of N-acetylmuramoyl-L-alanine Amidase from Staphylococcus aureus.


X-ray diffraction data for the 1.75 Angstrom Crystal Structure of Superantigen-like Protein, Exotoxin from Staphylococcus aureus, in Complex with Sialyl-LewisX.


X-ray diffraction data for the 1.90 Angstrom Resolution Crystal Structure of Glutathione Reductase from Streptococcus pyogenes in Complex with FAD.


X-ray diffraction data for the 2.25 Angstrom Resolution Crystal Structure of Fatty-Acid-CoA Ligase (FadD32) from Mycobacterium smegmatis in complex with Inhibitor 5'-O-[(11-phenoxyundecanoyl)sulfamoyl]adenosine.


X-ray diffraction data for the Structure of spermidine N-acetyltransferase SpeG from Vibrio cholerae


X-ray diffraction data for the 1.9 Angstrom resolution crystal structure of Se-methionine hypothetical protein SAOUHSC_02783 from Staphylococcus aureus


X-ray diffraction data for the Structure of the Salmonella typhimurium nfnB dihydropteridine reductase

First author:
S.M. Anderson
Gene name: nfnB
Resolution: 2.40 Å
R/Rfree: 0.18/0.22
Gene name: nfnB
Resolution: 2.40 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of glutamate racemase from Francisella tularensis subsp. tularensis SCHU S4 in complex with D-glutamate.

First author:
E.V. Filippova
Gene name: murI
Resolution: 1.65 Å
R/Rfree: 0.16/0.19
Gene name: murI
Resolution: 1.65 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Structure of the Bacillus anthracis tetrahydropicolinate succinyltransferase
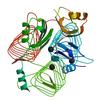
First author:
S.M. Anderson
Resolution: 1.70 Å
R/Rfree: 0.19/0.22
Resolution: 1.70 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the 1.84 Angstrom resolution crystal structure of 3-oxoacyl-(acyl carrier protein) synthase I (fabB) from Yersinia pestis CO92

First author:
A.S. Halavaty
Gene name: fabB
Resolution: 1.84 Å
R/Rfree: 0.16/0.19
Gene name: fabB
Resolution: 1.84 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of the yhdH oxidoreductase from Salmonella enterica in complex with NADP

First author:
S.M. Anderson
Gene name: yhdH
Resolution: 1.90 Å
R/Rfree: 0.15/0.19
Gene name: yhdH
Resolution: 1.90 Å
R/Rfree: 0.15/0.19
X-ray diffraction data for the Crystal Structure of a propionate kinase from Francisella tularensis subsp. tularensis SCHU S4

First author:
J.S. Brunzelle
Gene name: tdcD
Resolution: 1.98 Å
R/Rfree: 0.16/0.19
Gene name: tdcD
Resolution: 1.98 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the The crystal structure of kanamycin B dioxygenase (KanJ) from Streptomyces kanamyceticus in complex with nickel, sulfate, soaked with iodide


First author:
B. Mrugala
Resolution: 2.10 Å
R/Rfree: 0.17/0.20
Resolution: 2.10 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal Structure of the Sugar Binding Domain of LacI Family Protein from Klebsiella pneumoniae


X-ray diffraction data for the Structure of a putative aminopeptidase P from Bacillus anthracis

First author:
S.M. Anderson
Resolution: 2.89 Å
R/Rfree: 0.20/0.23
Resolution: 2.89 Å
R/Rfree: 0.20/0.23
X-ray diffraction data for the Crystal structure of the Bacillus anthracis acetyl-CoA acetyltransferase

First author:
S.M. Anderson
Resolution: 1.70 Å
R/Rfree: 0.15/0.17
Resolution: 1.70 Å
R/Rfree: 0.15/0.17
X-ray diffraction data for the Crystal structure of a putative uncharacterized protein from Mycobacterium tuberculosis


First author:
J. Abendroth
Resolution: 1.95 Å
R/Rfree: 0.18/0.22
Resolution: 1.95 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of the putative periplasmic solute-binding protein from Campylobacter jejuni


X-ray diffraction data for the 1.55 Angstrom Crystal Structure of N-acetylmuramic acid 6-phosphate Etherase from Yersinia enterocolitica.


X-ray diffraction data for the Cycloalternan-degrading enzyme from Trueperella pyogenes in complex with cycloalternan


X-ray diffraction data for the 1.85 Angstrom resolution crystal structure of an ABC transporter from Clostridium perfringens ATCC 13124

First author:
A.S. Halavaty
Resolution: 1.85 Å
R/Rfree: 0.16/0.19
Resolution: 1.85 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal Structure of a Phosphocarrier Protein HPr from Bacillus anthracis str. Ames

First author:
J.S. Brunzelle
Gene name: ptsH-2
Resolution: 1.15 Å
R/Rfree: 0.16/0.18
Gene name: ptsH-2
Resolution: 1.15 Å
R/Rfree: 0.16/0.18
X-ray diffraction data for the 1.05 Angstrom crystal structure of an amino acid ABC transporter substrate-binding protein AbpA from Streptococcus pneumoniae Canada MDR_19A bound to L-arginine

First author:
P.J. Stogios
Resolution: 1.05 Å
R/Rfree: 0.17/0.18
Resolution: 1.05 Å
R/Rfree: 0.17/0.18
X-ray diffraction data for the Crystal structure of apo transketolase from Francisella tularensis

First author:
S.M. Anderson
Gene name: tktA
Resolution: 1.60 Å
R/Rfree: 0.16/0.20
Gene name: tktA
Resolution: 1.60 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the 1.5 Angstrom Crystal Structure of Spermidine/Putrescine ABC Transporter Substrate-Binding Protein PotD from Streptococcus pneumoniae strain Canada MDR_19A in Complex with Calcium and HEPES

First author:
G. Minasov
Resolution: 1.50 Å
R/Rfree: 0.15/0.19
Resolution: 1.50 Å
R/Rfree: 0.15/0.19
X-ray diffraction data for the Structure of the YghA Oxidoreductase from Salmonella enterica

First author:
S.M. Anderson
Gene name: yghA
Resolution: 1.25 Å
R/Rfree: 0.13/0.15
Gene name: yghA
Resolution: 1.25 Å
R/Rfree: 0.13/0.15
X-ray diffraction data for the 1.9 Angstrom resolution crystal structure of a NAD synthetase (nadE) from Salmonella typhimurium LT2 in complex with NAD(+)

First author:
A.S. Halavaty
Gene name: nadE
Resolution: 1.90 Å
R/Rfree: 0.14/0.19
Gene name: nadE
Resolution: 1.90 Å
R/Rfree: 0.14/0.19
X-ray diffraction data for the X-ray Crystal Structure of a Putative Phosphate ABC Transporter Substrate-Binding Protein with Bound Phosphate from Clostridium perfringens


First author:
J.S. Brunzelle
Gene name: pstS
Resolution: 1.70 Å
R/Rfree: 0.15/0.19
Gene name: pstS
Resolution: 1.70 Å
R/Rfree: 0.15/0.19
X-ray diffraction data for the An X-ray Crystal Structure of a putative Bifunctional Phosphoribosylaminoimidazolecarboxamide Formyltransferase/IMP Cyclohydrolase

First author:
J.S. Brunzelle
Gene name: purH
Resolution: 2.28 Å
R/Rfree: 0.17/0.21
Gene name: purH
Resolution: 2.28 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of an aminoglycoside acetyltransferase meta-AAC0020 from an uncultured soil metagenomic sample in complex with trehalose

First author:
Z. Xu
Gene name: None
Resolution: 1.50 Å
R/Rfree: 0.16/0.18
Gene name: None
Resolution: 1.50 Å
R/Rfree: 0.16/0.18
X-ray diffraction data for the Crystal structure of vancomycin resistance D,D-dipeptidase/D,D-pentapeptidase VanXYc D59S mutant in complex with D-Alanine

First author:
P.J. Stogios
Gene name: vanXYc
Resolution: 2.25 Å
R/Rfree: 0.16/0.19
Gene name: vanXYc
Resolution: 2.25 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of the cytoplasmic domain of vancomycin resistance serine racemase VanTg

First author:
P.J. Stogios
Gene name: vanTG
Resolution: 2.02 Å
R/Rfree: 0.18/0.23
Gene name: vanTG
Resolution: 2.02 Å
R/Rfree: 0.18/0.23
X-ray diffraction data for the Crystal structure of serine hydroxymethyltransferase from Campylobacter jejuni

First author:
S.M. Anderson
Gene name: glyA
Resolution: 1.80 Å
R/Rfree: 0.16/0.20
Gene name: glyA
Resolution: 1.80 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the 2.4 Angstrom Resolution Crystal Structure of Putative Sugar Kinase from Campylobacter jejuni.

First author:
G. Minasov
Resolution: 2.40 Å
R/Rfree: 0.18/0.23
Resolution: 2.40 Å
R/Rfree: 0.18/0.23
X-ray diffraction data for the Crystal structure of Bacillus anthracis pyrrolidone-carboxylate peptidase, pcP

First author:
S.M. Anderson
Gene name: pcP
Resolution: 2.00 Å
R/Rfree: 0.18/0.22
Gene name: pcP
Resolution: 2.00 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystals structure of a Bacillus anthracis aminotransferase

First author:
S.M. Anderson
Resolution: 2.05 Å
R/Rfree: 0.16/0.19
Resolution: 2.05 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of rifampin phosphotransferase RPH-Lm from Listeria monocytogenes in complex with rifampin

First author:
P.J. Stogios
Resolution: 2.70 Å
R/Rfree: 0.22/0.26
Resolution: 2.70 Å
R/Rfree: 0.22/0.26
X-ray diffraction data for the 1.1 Angstrom Crystal Structure of Putative Modulator of Drug Activity (MdaB) from Yersinia pestis CO92.

First author:
G. Minasov
Gene name: mdaB
Resolution: 1.10 Å
R/Rfree: 0.10/0.12
Gene name: mdaB
Resolution: 1.10 Å
R/Rfree: 0.10/0.12
X-ray diffraction data for the Structure of UDP-N-acetylmuramoylalanyl-D-glutamyl-2,6-diaminopimelate--D-alanyl-D-alanyl ligase from Acinetobacter baumannii


First author:
Abendroth Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.85 Å
R/Rfree: 0.17/0.20
Resolution: 1.85 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of a dihydroorotase from Burkholderia cenocepacia J2315


First author:
C.M. Lukacs
Resolution: 1.80 Å
R/Rfree: 0.16/0.19
Resolution: 1.80 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the X-ray crystal structure of a putative D-amino acid aminotransferase from Burkholderia cenocepacia


First author:
J.W. Fairman
Resolution: 1.80 Å
R/Rfree: 0.17/0.21
Resolution: 1.80 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of Bovine Serum Albumin


First author:
K.A. Majorek
Resolution: 2.70 Å
R/Rfree: 0.20/0.26
Resolution: 2.70 Å
R/Rfree: 0.20/0.26
X-ray diffraction data for the 1.7 Angstrom resolution crystal structure of an acyl carrier protein S-malonyltransferase from Vibrio cholerae O1 biovar eltor str. N16961

First author:
A.S. Halavaty
Resolution: 1.70 Å
R/Rfree: 0.15/0.19
Resolution: 1.70 Å
R/Rfree: 0.15/0.19
X-ray diffraction data for the Structure of the N-terminal domain of lipoprotein-releasing system transmembrane protein LolE from Acinetobacter baumannii


First author:
J. Abendroth
Resolution: 2.35 Å
R/Rfree: 0.18/0.22
Resolution: 2.35 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of a Peptide Deformylase from Burkholderia multivorans


First author:
K.T. Potts
Resolution: 1.95 Å
R/Rfree: 0.17/0.21
Resolution: 1.95 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the 1.45 Angstrom Resolution Crystal Structure of PDZ domain of Carboxy-Terminal Protease from Vibrio cholerae in Complex with Peptide.


X-ray diffraction data for the Crystal structure of peptidase B from Yersinia pestis CO92 at 2.75 A resolution


X-ray diffraction data for the X-ray crystal structure of a putative UDP-4-amino-4-deoxy-l-arabinose--oxoglutarate aminotransferase from Burkholderia cenocepacia


First author:
J.W. Fairman
Resolution: 1.60 Å
R/Rfree: 0.11/0.15
Resolution: 1.60 Å
R/Rfree: 0.11/0.15
X-ray diffraction data for the Catalytic domain of LPMO Lmo2467 from Listeria monocytogenes


X-ray diffraction data for the 2.2 Angstrom Crystal Structure of ABC Transporter Substrate Binding Protein CtaP (Lmo0135) from Listeria monocytogenes.


X-ray diffraction data for the 2.6 Angstrom Resolution Crystal Structure of Stage II Sporulation Protein D (SpoIID) from Clostridium difficile in Complex with Triacetylchitotriose


X-ray diffraction data for the 1.9 Angstrom Crystal Structure of NS5 Methyl Transferase from Dengue Virus 1 in Complex with S-Adenosylmethionine and Beta-D-Fructopyranose.

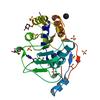
X-ray diffraction data for the 1.78 Angstrom Crystal Structure of the Salmonella enterica 3-Dehydroquinate Dehydratase (aroD) in Complex with Quinate


X-ray diffraction data for the Cycloalternan-degrading enzyme from Trueperella pyogenes in complex with covalent intermediate


X-ray diffraction data for the Crystal structure of Oxidoreductase, 2OG-Fe(II) oxygenase family, from Burkholderia pseudomallei


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 2.05 Å
R/Rfree: 0.21/0.24
Resolution: 2.05 Å
R/Rfree: 0.21/0.24
X-ray diffraction data for the 1.67 Angstrom Resolution Crystal Structure of Murein-DD-endopeptidase from Yersinia enterocolitica.


X-ray diffraction data for the Crystal structure of spermidine/spermine N-acetyltransferase SpeG from Escherichia coli in complex with tris(hydroxymethyl)aminomethane.


X-ray diffraction data for the Structure of a GNAT superfamily PA3944 acetyltransferase in complex with AcCoA


First author:
M.P. Czub
Resolution: 1.80 Å
R/Rfree: 0.21/0.24
Resolution: 1.80 Å
R/Rfree: 0.21/0.24
X-ray diffraction data for the Crystal Structure of Peptidylprolyl Isomerase PrsA from Streptococcus mutans.


First author:
G. Minasov
Gene name: prtM
Resolution: 3.15 Å
R/Rfree: 0.27/0.31
Gene name: prtM
Resolution: 3.15 Å
R/Rfree: 0.27/0.31
X-ray diffraction data for the Aminopeptidase N (pepN) from Francisella tularensis subsp. tularensis SCHU S4

First author:
D. Borek
Gene name: pepN
Resolution: 2.51 Å
R/Rfree: 0.19/0.26
Gene name: pepN
Resolution: 2.51 Å
R/Rfree: 0.19/0.26
X-ray diffraction data for the Crystal structure of the competence-damaged protein (CinA) superfamily protein ECK1530/EC0983 from Escherichia coli

First author:
P.J. Stogios
Gene name: ydeJ
Resolution: 1.91 Å
R/Rfree: 0.21/0.25
Gene name: ydeJ
Resolution: 1.91 Å
R/Rfree: 0.21/0.25
X-ray diffraction data for the 2.15 Angstrom Crystal Structure of S-adenosylhomocysteinase from Cryptosporidium parvum in Complex with D-Eritadenine and NAD

First author:
G. Minasov
Resolution: 2.15 Å
R/Rfree: 0.14/0.18
Resolution: 2.15 Å
R/Rfree: 0.14/0.18
X-ray diffraction data for the Crystal structure of protein of unknown function ORF19 from Vibrio cholerae O1 PICI-like element, C57S I109M mutant

First author:
P.J. Stogios
Resolution: 3.30 Å
R/Rfree: 0.24/0.29
Resolution: 3.30 Å
R/Rfree: 0.24/0.29
X-ray diffraction data for the 1.76 Angstrom Crystal Structure of GTP-binding Protein Der from Coxiella burnetii in Complex with GDP.

First author:
G. Minasov
Resolution: 1.76 Å
R/Rfree: 0.17/0.20
Resolution: 1.76 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of the ribosomal-protein-S18-alanine N-acetyltransferase from Escherichia coli

First author:
E.V. Filippova
Gene name: rimI
Resolution: 1.35 Å
R/Rfree: 0.16/0.21
Gene name: rimI
Resolution: 1.35 Å
R/Rfree: 0.16/0.21
X-ray diffraction data for the 1.76A X-ray Crystal Structure of a Putative Phenazine Biosynthesis PhzC/PhzF Protein from Clostridium difficile (strain 630)

First author:
J.S. Brunzelle
Resolution: 1.76 Å
R/Rfree: 0.16/0.19
Resolution: 1.76 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the 1.80 Angstrom resolution crystal structure of orotidine 5'-phosphate decarboxylase from Vibrio cholerae O1 biovar eltor str. N16961 in complex with uridine-5'-monophosphate (UMP)

First author:
A.S. Halavaty
Resolution: 1.80 Å
R/Rfree: 0.15/0.18
Resolution: 1.80 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the 1.75 Angstrom resolution crystal structure of an orotate phosphoribosyltransferase from Bacillus anthracis str. 'Ames Ancestor'

First author:
A.S. Halavaty
Gene name: pyrE
Resolution: 1.75 Å
R/Rfree: 0.18/0.22
Gene name: pyrE
Resolution: 1.75 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Structure of an autocleavage-inactive mutant of the cytoplasmic domain of CT091, the YscU homologue of Chlamydia trachomatis

First author:
A.U. Singer
Gene name: yscU
Resolution: 2.10 Å
R/Rfree: 0.19/0.22
Gene name: yscU
Resolution: 2.10 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the 2.9 Angstrom Crystal Structure of Putative Exotoxin 3 from Staphylococcus aureus.
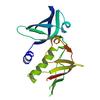
First author:
G. Minasov
Resolution: 2.90 Å
R/Rfree: 0.23/0.28
Resolution: 2.90 Å
R/Rfree: 0.23/0.28
X-ray diffraction data for the 2.6 Angstrom Crystal Structure of UDP-N-acetylglucosamine 1-carboxyvinyltransferase 1 (MurA1) from Bacillus anthracis

First author:
G. Minasov
Resolution: 2.60 Å
R/Rfree: 0.18/0.25
Resolution: 2.60 Å
R/Rfree: 0.18/0.25
X-ray diffraction data for the 1.4 Angstrom Crystal Structure of ABC Transporter Glutathione-Binding Protein GshT from Streptococcus pneumoniae strain Canada MDR_19A in Complex with Glutathione

First author:
G. Minasov
Resolution: 1.40 Å
R/Rfree: 0.17/0.19
Resolution: 1.40 Å
R/Rfree: 0.17/0.19
X-ray diffraction data for the Crystal structure of the aminoglycoside phosphotransferase APH(3')-Ia, with substrate kanamycin and small molecule inhibitor 1-NM-PP1

First author:
P.J. Stogios
Gene name: aphA1
Resolution: 1.88 Å
R/Rfree: 0.17/0.22
Gene name: aphA1
Resolution: 1.88 Å
R/Rfree: 0.17/0.22
X-ray diffraction data for the 1.8 Angstrom Resolution Crystal Structure of Cytosol Aminopeptidase from Coxiella burnetii

First author:
G. Minasov
Resolution: 1.80 Å
R/Rfree: 0.15/0.18
Resolution: 1.80 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Potassium transporter peripheral membrane component (trkA) from Vibrio vulnificus

First author:
E.V. Filippova
Gene name: trkA
Resolution: 2.09 Å
R/Rfree: 0.18/0.23
Gene name: trkA
Resolution: 2.09 Å
R/Rfree: 0.18/0.23
X-ray diffraction data for the Crystal Structure of a Putative Succinate-Semialdehyde Dehydrogenase from Salmonella typhimurium LT2 with bound NAD

First author:
J.S. Brunzelle
Gene name: yneI
Resolution: 1.90 Å
R/Rfree: 0.16/0.19
Gene name: yneI
Resolution: 1.90 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the 2.3 Angstrom Crystal Structure of a Glucose-1-phosphate Thymidylyltransferase from Bacillus anthracis in Complex with Thymidine-5-diphospho-alpha-D-glucose and Pyrophosphate

First author:
G. Minasov
Resolution: 2.30 Å
R/Rfree: 0.15/0.19
Resolution: 2.30 Å
R/Rfree: 0.15/0.19
X-ray diffraction data for the 1.43 Angstrom Resolution Crystal Structure of Triosephosphate Isomerase (tpiA) from Escherichia coli in Complex with Acetyl Phosphate.

First author:
G. Minasov
Gene name: tpiA
Resolution: 1.43 Å
R/Rfree: 0.13/0.16
Gene name: tpiA
Resolution: 1.43 Å
R/Rfree: 0.13/0.16
X-ray diffraction data for the 1.37 Angstrom resolution crystal structure of apo-form of a putative deoxyribose-phosphate aldolase from Toxoplasma gondii ME49

First author:
A.S. Halavaty
Resolution: 1.37 Å
R/Rfree: 0.15/0.18
Resolution: 1.37 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the 1.5 Angstrom crystal structure of soluble domain of membrane-anchored thioredoxin family protein from Streptococcus pneumoniae strain Canada MDR_19A

First author:
Z. Wawrzak
Resolution: 1.51 Å
R/Rfree: 0.17/0.20
Resolution: 1.51 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of an amino acid ABC transporter substrate-binding protein from Streptococcus pneumoniae Canada MDR_19A bound to L-arginine, form 1

First author:
P.J. Stogios
Resolution: 1.90 Å
R/Rfree: 0.16/0.20
Resolution: 1.90 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the 1.5 Angstrom Resolution Crystal Structure of K135M Mutant of Transaldolase B (TalA) from Francisella tularensis in Complex with Sedoheptulose 7-phosphate.

First author:
G. Minasov
Gene name: talA
Resolution: 1.50 Å
R/Rfree: 0.14/0.16
Gene name: talA
Resolution: 1.50 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the 2.75 Angstrom Crystal Structure of Hypothetical Protein lmo2686 from Listeria monocytogenes EGD-e

First author:
G. Minasov
Gene name: -
Resolution: 2.75 Å
R/Rfree: 0.20/0.24
Gene name: -
Resolution: 2.75 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the NH3-dependent NAD synthetase from Campylobacter jejuni subsp. jejuni NCTC 11168 in complex with the nitrate ion

First author:
E.V. Filippova
Gene name: nadE
Resolution: 2.74 Å
R/Rfree: 0.21/0.24
Gene name: nadE
Resolution: 2.74 Å
R/Rfree: 0.21/0.24
X-ray diffraction data for the Crystal structure of the aminoglycoside phosphotransferase APH(3')-Ia, with substrate kanamycin and small molecule inhibitor pyrazolopyrimidine PP2

First author:
P.J. Stogios
Gene name: aphA1
Resolution: 1.98 Å
R/Rfree: 0.16/0.22
Gene name: aphA1
Resolution: 1.98 Å
R/Rfree: 0.16/0.22
X-ray diffraction data for the 2.60 Angstrom resolution crystal structure of betaine aldehyde dehydrogenase (betB) H448F/Y450L double mutant from Staphylococcus aureus in complex with NAD+ and BME-free Cys289

First author:
A.S. Halavaty
Gene name: betB
Resolution: 2.60 Å
R/Rfree: 0.16/0.23
Gene name: betB
Resolution: 2.60 Å
R/Rfree: 0.16/0.23
X-ray diffraction data for the 2.85 Angstrom resolution crystal structure of betaine aldehyde dehydrogenase (betB) H448F/P449M double mutant from Staphylococcus aureus in complex with NAD+ and BME-free Cys289

First author:
A.S. Halavaty
Gene name: betB
Resolution: 2.85 Å
R/Rfree: 0.16/0.21
Gene name: betB
Resolution: 2.85 Å
R/Rfree: 0.16/0.21
X-ray diffraction data for the Crystal Structure of K170M Mutant of Type I 3-Dehydroquinate Dehydratase (aroD) from Salmonella typhimurium LT2 in Non-Covalent Complex with Dehydroquinate.

First author:
G. Minasov
Gene name: aroD
Resolution: 1.60 Å
R/Rfree: 0.16/0.19
Gene name: aroD
Resolution: 1.60 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of anabolic ornithine carbamoyltransferase from Vibrio vulnificus in complex with citrulline and inorganic phosphate

First author:
I.G. Shabalin
Resolution: 2.08 Å
R/Rfree: 0.16/0.19
Resolution: 2.08 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of anabolic ornithine carbamoyltransferase from Vibrio vulnificus in complex with carbamoylphosphate and arginine

First author:
I.G. Shabalin
Resolution: 1.85 Å
R/Rfree: 0.15/0.19
Resolution: 1.85 Å
R/Rfree: 0.15/0.19
X-ray diffraction data for the Crystal structure of iron uptake ABC transporter substrate-binding protein PiaA from Streptococcus pneumoniae Canada MDR_19A bound to Bis-tris propane

First author:
P.J. Stogios
Resolution: 1.66 Å
R/Rfree: 0.17/0.19
Resolution: 1.66 Å
R/Rfree: 0.17/0.19
X-ray diffraction data for the Crystal structure of a putative Cas1 enzyme from Vibrio phage ICP1

First author:
P.J. Stogios
Gene name: cas1
Resolution: 2.13 Å
R/Rfree: 0.17/0.21
Gene name: cas1
Resolution: 2.13 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal Structure of Type I 3-Dehydroquinate Dehydratase (aroD) from Salmonella typhimurium with close loop conformation.

First author:
G. Minasov
Gene name: aroD
Resolution: 1.90 Å
R/Rfree: 0.15/0.20
Gene name: aroD
Resolution: 1.90 Å
R/Rfree: 0.15/0.20
X-ray diffraction data for the 2.09 Angstrom resolution structure of a hypoxanthine-guanine phosphoribosyltransferase (hpt-1) from Bacillus anthracis str. 'Ames Ancestor' in complex with GMP

First author:
A.S. Halavaty
Gene name: hpt-1
Resolution: 2.09 Å
R/Rfree: 0.20/0.24
Gene name: hpt-1
Resolution: 2.09 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the 1.70 Angstrom resolution crystal structure of outer-membrane lipoprotein carrier protein (lolA) from Yersinia pestis CO92

First author:
A.S. Halavaty
Gene name: lolA
Resolution: 1.70 Å
R/Rfree: 0.20/0.24
Gene name: lolA
Resolution: 1.70 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the Crystal structure of spermidine N-acetyltransferase from Vibrio cholerae in complex with spermine

First author:
E.V. Filippova
Resolution: 1.85 Å
R/Rfree: 0.15/0.18
Resolution: 1.85 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the 1.95 Angstrom Resolution Crystal Structure of Epidermin Leader Peptide Processing Serine Protease (EpiP) S393A Mutant from Staphylococcus aureus

First author:
G. Minasov
Gene name: epiP
Resolution: 1.95 Å
R/Rfree: 0.17/0.19
Gene name: epiP
Resolution: 1.95 Å
R/Rfree: 0.17/0.19
X-ray diffraction data for the 2.4 Angstrom Crystal Structure of Dihydroorotase (pyrC) from Campylobacter jejuni.

First author:
G. Minasov
Gene name: pyrC
Resolution: 2.40 Å
R/Rfree: 0.19/0.25
Gene name: pyrC
Resolution: 2.40 Å
R/Rfree: 0.19/0.25
X-ray diffraction data for the 1.8 Angstrom resolution crystal structure of dihydroorotase (pyrC) from Salmonella enterica subsp. enterica serovar Typhimurium str. LT2

First author:
G. Minasov
Gene name: pyrC
Resolution: 1.80 Å
R/Rfree: 0.16/0.20
Gene name: pyrC
Resolution: 1.80 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the Crystal Structure of the 3-Dehydroquinate Dehydratase (aroD) from Salmonella typhimurium LT2 with Citrate Bound to the Active Site.

First author:
G. Minasov
Gene name: aroD
Resolution: 1.65 Å
R/Rfree: 0.16/0.20
Gene name: aroD
Resolution: 1.65 Å
R/Rfree: 0.16/0.20