6726 results
X-ray diffraction data for the CRYSTAL STRUCTURE OF A UBIQUINONE/MENAQUINONE BIOSYNTHESIS METHYLTRANSFERASE-RELATED PROTEIN (TM1389) FROM THERMOTOGA MARITIMA MSB8 AT 2.35 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.35 Å
R/Rfree: 0.17/0.20
Resolution: 2.35 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of Dephospho-CoA kinase (EC 2.7.1.24) (Dephosphocoenzyme A kinase) (tm1387) from THERMOTOGA MARITIMA at 2.60 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.60 Å
R/Rfree: 0.19/0.25
Resolution: 2.60 Å
R/Rfree: 0.19/0.25
X-ray diffraction data for the Crystal structure of Glucose-6-phosphate isomerase (EC 5.3.1.9) (TM1385) from Thermotoga maritima at 1.82 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.82 Å
R/Rfree: 0.20/0.24
Resolution: 1.82 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the Crystal structure of a predicted rna methyltransferase (tm1380) from thermotoga maritima msb8 at 2.12 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.12 Å
R/Rfree: 0.17/0.22
Resolution: 2.12 Å
R/Rfree: 0.17/0.22
X-ray diffraction data for the CRYSTAL STRUCTURE OF an atypical cyclophilin (peptidylprolyl cis-trans isomerase) (TM1367) FROM THERMOTOGA MARITIMA AT 1.90 A RESOLUTION

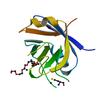
First author:
K.K. Jin
Resolution: 1.90 Å
R/Rfree: 0.17/0.21
Resolution: 1.90 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of Glutamyl-tRNA synthetase 1 (EC 6.1.1.17) (Glutamate-tRNA ligase 1) (GluRS 1) (TM1351) from Thermotoga maritima at 2.5 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.34 Å
R/Rfree: 0.23/0.27
Resolution: 2.34 Å
R/Rfree: 0.23/0.27
X-ray diffraction data for the Crystal structure of Inosine-5'-monophosphate dehydrogenase (TM1347) from THERMOTOGA MARITIMA at 2.18 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.18 Å
R/Rfree: 0.22/0.26
Resolution: 2.18 Å
R/Rfree: 0.22/0.26
X-ray diffraction data for the Crystal structure of SAM-dependent methyltransferase, possible histamine N-methyltransferase (TM1293) from Thermotoga maritima at 2.20 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.20 Å
R/Rfree: 0.19/0.25
Resolution: 2.20 Å
R/Rfree: 0.19/0.25
X-ray diffraction data for the Crystal structure of a predicted oxalate decarboxylase (tm1287) from thermotoga maritima at 1.95 A resolution


First author:
R. Schwarzenbacher
Resolution: 1.95 Å
R/Rfree: 0.16/0.22
Resolution: 1.95 Å
R/Rfree: 0.16/0.22
X-ray diffraction data for the Crystal structure of Glutamyl-tRNA(Gln) amidotransferase subunit A (tm1272) from THERMOTOGA MARITIMA at 1.80 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.80 Å
R/Rfree: 0.18/0.20
Resolution: 1.80 Å
R/Rfree: 0.18/0.20
X-ray diffraction data for the Crystal structure of Aspartate aminotransferase (TM1255) from Thermotoga maritima at 1.90 A resolution


First author:
R. Schwarzenbacher
Resolution: 1.90 Å
R/Rfree: 0.16/0.20
Resolution: 1.90 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the Crystal structure of Phosphoribosylamine--glycine ligase (TM1250) from Thermotoga maritima at 2.30 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.30 Å
R/Rfree: 0.21/0.26
Resolution: 2.30 Å
R/Rfree: 0.21/0.26
X-ray diffraction data for the Crystal structure of Phosphoribosylaminoimidazolecarboxamide formyltransferase / IMP cyclohydrolase (TM1249) from THERMOTOGA MARITIMA at 1.88 A resolution


First author:
H.L. Axelrod
Resolution: 1.88 Å
R/Rfree: 0.16/0.20
Resolution: 1.88 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the Crystal structure of Phosphoribosylformylglycinamidine synthase II (TM1246) from Thermotoga maritima at 2.15 A resolution


First author:
I.I. Mathews
Resolution: 2.15 Å
R/Rfree: 0.19/0.25
Resolution: 2.15 Å
R/Rfree: 0.19/0.25
X-ray diffraction data for the Crystal structure of Phosphoribosylformylglycinamidine synthase, purS subunit (EC 6.3.5.3) (TM1244) from Thermotoga maritima at 1.90 A resolution


First author:
I.I. Mathews
Resolution: 1.90 Å
R/Rfree: 0.18/0.24
Resolution: 1.90 Å
R/Rfree: 0.18/0.24
X-ray diffraction data for the Crystal structure of a predicted glycosidase (tm1225) from thermotoga maritima msb8 at 2.10 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.10 Å
R/Rfree: 0.16/0.20
Resolution: 2.10 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the Crystal structure of N-acetylglucosamine kinase (TM1224) from Thermotoga maritima at 2.46 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.46 Å
R/Rfree: 0.22/0.27
Resolution: 2.46 Å
R/Rfree: 0.22/0.27
X-ray diffraction data for the Crystal structure of Oligopeptide ABC transporter, periplasmic oligopeptide-binding (TM1223) from THERMOTOGA MARITIMA at 1.73 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.73 Å
R/Rfree: 0.15/0.18
Resolution: 1.73 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of Alpha-galactosidase (EC 3.2.1.22) (Melibiase) (tm1192) from Thermotoga maritima at 2.34 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.34 Å
R/Rfree: 0.16/0.21
Resolution: 2.34 Å
R/Rfree: 0.16/0.21
X-ray diffraction data for the Crystal structure of Methylglyoxal synthase (TM1185) from Thermotoga maritima at 2.06 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.06 Å
R/Rfree: 0.17/0.22
Resolution: 2.06 Å
R/Rfree: 0.17/0.22
X-ray diffraction data for the Crystal structure of 3-oxoacyl-(acyl carrier protein) reductase (TM1169) from Thermotoga maritima at 2.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.50 Å
R/Rfree: 0.19/0.25
Resolution: 2.50 Å
R/Rfree: 0.19/0.25
X-ray diffraction data for the Crystal structure of 6-phosphogluconolactonase (TM1154) from Thermotoga maritima at 1.70A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.55 Å
R/Rfree: 0.17/0.20
Resolution: 1.55 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of a Leucine binding protein LivK (TM1135) from Thermotoga maritima MSB8 at 1.90 A resolution


First author:
Joint center for structural genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.15/0.19
Resolution: 1.90 Å
R/Rfree: 0.15/0.19
X-ray diffraction data for the Crystal structure of a putative aminotransferase (tm1131) from thermotoga maritima msb8 at 1.82 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.82 Å
R/Rfree: 0.18/0.20
Resolution: 1.82 Å
R/Rfree: 0.18/0.20
X-ray diffraction data for the Crystal structure of Ferritin (TM1128) from Thermotoga maritima at 2.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.18/0.22
Resolution: 2.00 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of a duf861 family protein (tm1112) from thermotoga maritima at 1.83 A resolution

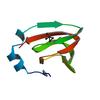
First author:
D. McMullan
Resolution: 1.83 Å
R/Rfree: 0.17/0.22
Resolution: 1.83 Å
R/Rfree: 0.17/0.22
X-ray diffraction data for the Crystal structure of Ribonuclease III (TM1102) from Thermotoga maritima at 2.0 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.20/0.23
Resolution: 2.00 Å
R/Rfree: 0.20/0.23
X-ray diffraction data for the Crystal structure of Ornithine carbamoyltransferase (TM1097) from Thermotoga maritima at 2.25 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.25 Å
R/Rfree: 0.19/0.22
Resolution: 2.25 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE TRANSPORT PROTEIN (TM1088A) FROM THERMOTOGA MARITIMA AT 1.50 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.50 Å
R/Rfree: 0.15/0.18
Resolution: 1.50 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the CRYSTAL STRUCTURE OF AN ARCHEASE, POSSIBLE CHAPERONE (TM1083) FROM THERMOTOGA MARITIMA AT 2.0 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.21/0.27
Resolution: 2.00 Å
R/Rfree: 0.21/0.27
X-ray diffraction data for the Crystal structure of a putative anti-sigma factor antagonist (tm1081) from thermotoga maritima at 1.59 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.59 Å
R/Rfree: 0.18/0.21
Resolution: 1.59 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of a ribose 5-phosphate isomerase rpib (tm1080) from thermotoga maritima at 1.90 A resolution


First author:
Q. Xu
Resolution: 1.90 Å
R/Rfree: 0.12/0.14
Resolution: 1.90 Å
R/Rfree: 0.12/0.14
X-ray diffraction data for the Crystal structure of Periplasmic divalent cation tolerance protein (TM1056) from Thermotoga maritima at 1.95 A resolution
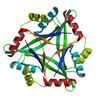
First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.95 Å
R/Rfree: 0.16/0.22
Resolution: 1.95 Å
R/Rfree: 0.16/0.22
X-ray diffraction data for the Crystal structure of Endoglucanase (TM1050) from THERMOTOGA MARITIMA at 1.40 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.40 Å
R/Rfree: 0.15/0.18
Resolution: 1.40 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of Endoglucanase (tm1049) from THERMOTOGA MARITIMA at 2.01 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.01 Å
R/Rfree: 0.20/0.24
Resolution: 2.01 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the Crystal structure of Histidinol-phosphate aminotransferase (EC 2.6.1.9) (Imidazole acetol-phosphate transferase) (tm1040) from Thermotoga maritima at 2.40 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.40 Å
R/Rfree: 0.18/0.24
Resolution: 2.40 Å
R/Rfree: 0.18/0.24
X-ray diffraction data for the Crystal structure of Imidazole glycerol phosphate synthase subunit hisF (EC 4.1.3.-) (tm1036) from Thermotoga maritima at 1.64 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.64 Å
R/Rfree: 0.17/0.19
Resolution: 1.64 Å
R/Rfree: 0.17/0.19
X-ray diffraction data for the Crystal structure of Transcriptional regulator, TETR family (tm1030) from Thermotoga maritima at 2.30 A resolution


First author:
L. Premkumar
Resolution: 2.30 Å
R/Rfree: 0.20/0.26
Resolution: 2.30 Å
R/Rfree: 0.20/0.26
X-ray diffraction data for the Crystal structure of a putative nucleotidyltransferase (tm1012) from Thermotoga maritima at 1.35 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.20 Å
R/Rfree: 0.14/0.17
Resolution: 1.20 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of a putative nucleotidyltransferase (tm1012) from Thermotoga maritima at 1.60 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.60 Å
R/Rfree: 0.15/0.18
Resolution: 1.60 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of a cupin-like protein (tm1010) from thermotoga maritima msb8 at 1.90 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.16/0.22
Resolution: 1.90 Å
R/Rfree: 0.16/0.22
X-ray diffraction data for the Crystal structure of 2,5-diketo-D-gluconic acid reductase (TM1009) from Thermotoga maritima at 2.40 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.40 Å
R/Rfree: 0.15/0.20
Resolution: 2.40 Å
R/Rfree: 0.15/0.20
X-ray diffraction data for the Crystal structure of a lema protein (tm0961) from thermotoga maritima msb8 at 2.28 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.28 Å
R/Rfree: 0.22/0.27
Resolution: 2.28 Å
R/Rfree: 0.22/0.27
X-ray diffraction data for the Crystal structure of Ribokinase (TM0960) from Thermotoga maritima at 2.15 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.15 Å
R/Rfree: 0.19/0.24
Resolution: 2.15 Å
R/Rfree: 0.19/0.24
X-ray diffraction data for the Crystal structure of an ob-fold protein (tm0957) from thermotoga maritima msb8 at 1.90 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.25 Å
R/Rfree: 0.19/0.25
Resolution: 2.25 Å
R/Rfree: 0.19/0.25
X-ray diffraction data for the Crystal structure of Metal-dependent hydrolase of cytosinedemaniase/chlorohydrolase family (TM0936) from Thermotoga maritima at 1.9 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.18/0.20
Resolution: 1.90 Å
R/Rfree: 0.18/0.20
X-ray diffraction data for the Crystal structure of a cbs domain-containing protein (tm0935) from thermotoga maritima at 1.87 A resolution


First author:
M.D. Miller
Resolution: 1.87 Å
R/Rfree: 0.19/0.26
Resolution: 1.87 Å
R/Rfree: 0.19/0.26
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE CARBOHYDRATE KINASE (TM0922) FROM THERMOTOGA MARITIMA MSB8 AT 2.25 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.27 Å
R/Rfree: 0.16/0.22
Resolution: 2.27 Å
R/Rfree: 0.16/0.22
X-ray diffraction data for the Crystal structure of Alcohol dehydrogenase, iron-containing (TM0920) from Thermotoga maritima at 1.30 A resolution


First author:
R. Schwarzenbacher
Resolution: 1.30 Å
R/Rfree: 0.14/0.17
Resolution: 1.30 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of Hydroperoxide resistance protein OsmC (TM0919) from Thermotoga maritima at 1.80 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.80 Å
R/Rfree: 0.17/0.22
Resolution: 1.80 Å
R/Rfree: 0.17/0.22
X-ray diffraction data for the Crystal structure of Ribonuclease HII (EC 3.1.26.4) (RNase HII) (tm0915) from THERMOTOGA MARITIMA at 1.74 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.74 Å
R/Rfree: 0.18/0.22
Resolution: 1.74 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of a putative metallo-beta-lactamase (tm0894) from Thermotoga maritima at 2.10 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.10 Å
R/Rfree: 0.16/0.18
Resolution: 2.10 Å
R/Rfree: 0.16/0.18
X-ray diffraction data for the CRYSTAL STRUCTURE OF A CBS DOMAIN PAIR/ACT DOMAIN PROTEIN (TM0892) FROM THERMOTOGA MARITIMA AT 1.70 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.25/0.27
Resolution: 1.70 Å
R/Rfree: 0.25/0.27
X-ray diffraction data for the Crystal structure of Homoserine O-succinyltransferase (EC 2.3.1.46) (Homoserine O-transsuccinylase) (HTS) (tm0881) from THERMOTOGA MARITIMA at 2.52 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.52 Å
R/Rfree: 0.19/0.27
Resolution: 2.52 Å
R/Rfree: 0.19/0.27
X-ray diffraction data for the Crystal structure of an orphan protein (TM0875) from Thermotoga maritima at 2.00 A resolution


First author:
C. Bakolitsa
Resolution: 2.00 Å
R/Rfree: 0.18/0.24
Resolution: 2.00 Å
R/Rfree: 0.18/0.24
X-ray diffraction data for the Crystal structure of Glycoprotein endopeptidase (tm0874) from THERMOTOGA MARITIMA at 2.50 A resolution


First author:
Q. Xu
Resolution: 2.50 Å
R/Rfree: 0.19/0.24
Resolution: 2.50 Å
R/Rfree: 0.19/0.24
X-ray diffraction data for the Crystal structure of a putative branched-chain amino acid aminotransferase (TM0831) from Thermotoga maritima at 2.15 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.15 Å
R/Rfree: 0.17/0.20
Resolution: 2.15 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of Possible 1-phosphofructokinase (EC 2.7.1.56) (tm0828) from Thermotoga Maritima at 2.46 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.46 Å
R/Rfree: 0.19/0.24
Resolution: 2.46 Å
R/Rfree: 0.19/0.24
X-ray diffraction data for the Crystal structure of NADH-dependent butanol dehydrogenase A (TM0820) from Thermotoga maritima at 1.78 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.78 Å
R/Rfree: 0.14/0.19
Resolution: 1.78 Å
R/Rfree: 0.14/0.19
X-ray diffraction data for the Crystal structure of a marr-like transcriptional regulator (tm0816) from thermotoga maritima at 2.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.30 Å
R/Rfree: 0.23/0.26
Resolution: 2.30 Å
R/Rfree: 0.23/0.26
X-ray diffraction data for the Crystal structure of N-acetylglucosamine-6-phosphate deacetylase (TM0814) from Thermotoga maritima at 2.5 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.50 Å
R/Rfree: 0.20/0.25
Resolution: 2.50 Å
R/Rfree: 0.20/0.25
X-ray diffraction data for the Crystal structure of putative nitroalkan dioxygenase (TM0800) from Thermotoga maritima at 2.71 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.71 Å
R/Rfree: 0.19/0.21
Resolution: 2.71 Å
R/Rfree: 0.19/0.21
X-ray diffraction data for the Crystal structure of a putative nucleotide binding protein (tm0796) from Thermotoga maritima at 2.67 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.67 Å
R/Rfree: 0.21/0.24
Resolution: 2.67 Å
R/Rfree: 0.21/0.24
X-ray diffraction data for the Crystal structure of a dna polymerase iii, gamma subunit-related protein (tm0771) from thermotoga maritima msb8 at 2.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.22/0.28
Resolution: 2.00 Å
R/Rfree: 0.22/0.28
X-ray diffraction data for the Crystal structure of Flavoprotein (TM0755) from Thermotoga maritima at 1.80 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.80 Å
R/Rfree: 0.15/0.18
Resolution: 1.80 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of Alpha-glucosidase (TM0752) from Thermotoga maritima at 2.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.50 Å
R/Rfree: 0.19/0.26
Resolution: 2.50 Å
R/Rfree: 0.19/0.26
X-ray diffraction data for the Crystal structure of SAM-dependent O-methyltransferase (TM0748) from Thermotoga maritima at 1.65 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.65 Å
R/Rfree: 0.16/0.19
Resolution: 1.65 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of Phosphopantetheine adenylyltransferase (TM0741) from Thermotoga maritima at 2.20 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.20 Å
R/Rfree: 0.17/0.23
Resolution: 2.20 Å
R/Rfree: 0.17/0.23
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE MODULATOR OF A DNA GYRASE (TM0727) FROM THERMOTOGA MARITIMA MSB8 AT 1.95 A RESOLUTION


First author:
C. Rife
Resolution: 1.95 Å
R/Rfree: 0.17/0.22
Resolution: 1.95 Å
R/Rfree: 0.17/0.22
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE THIAMIN PHOSPHATE SYNTHASE (TM0723) FROM THERMOTOGA MARITIMA MSB8 AT 1.52 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.52 Å
R/Rfree: 0.14/0.16
Resolution: 1.52 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the Crystal structure of propionyl-CoA carboxylase, beta subunit (TM0716) from THERMOTOGA MARITIMA at 2.30 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.30 Å
R/Rfree: 0.15/0.21
Resolution: 2.30 Å
R/Rfree: 0.15/0.21
X-ray diffraction data for the Crystal structure of a putative nucleic acid binding protein (tm0693) from thermotoga maritima at 2.28 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.28 Å
R/Rfree: 0.20/0.25
Resolution: 2.28 Å
R/Rfree: 0.20/0.25
X-ray diffraction data for the CRYSTAL STRUCTURE OF a putative nucleic acid binding protein (TM0693) FROM THERMOTOGA MARITIMA MSB8 AT 2.05 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.05 Å
R/Rfree: 0.19/0.25
Resolution: 2.05 Å
R/Rfree: 0.19/0.25
X-ray diffraction data for the CRYSTAL STRUCTURE OF A C-TERMINAL FRAGMENT OF THE PUTATIVE FLAGELLAR MOTOR SWITCH PROTEIN FLIN (TM0680) FROM THERMOTOGA MARITIMA AT 1.85 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.85 Å
R/Rfree: 0.17/0.21
Resolution: 1.85 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of TatD-related deoxyribonuclease (TM0667) from Thermotoga maritima at 1.8 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.80 Å
R/Rfree: 0.19/0.22
Resolution: 1.80 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of a putative superoxide reductase (TM0658) from THERMOTOGA MARITIMA at 1.10 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.10 Å
R/Rfree: 0.12/0.15
Resolution: 1.10 Å
R/Rfree: 0.12/0.15
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE SUPEROXIDE REDUCTASE (TM0658) FROM THERMOTOGA MARITIMA AT 2.00 A RESOLUTION

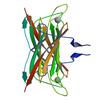
First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.15/0.20
Resolution: 2.00 Å
R/Rfree: 0.15/0.20
X-ray diffraction data for the Crystal structure of S-adenosylmethionine decarboxylase proenzyme (TM0655) from THERMOTOGA MARITIMA at 1.2 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.20 Å
Resolution: 1.20 Å
X-ray diffraction data for the Crystal structure of an hepn domain protein (tm0613) from thermotoga maritima at 1.75 A resolution


First author:
H. Erlandsen
Resolution: 1.75 Å
R/Rfree: 0.18/0.24
Resolution: 1.75 Å
R/Rfree: 0.18/0.24
X-ray diffraction data for the Crystal structure of single stranded DNA-binding protein (TM0604) from Thermotoga maritima at 2.60 A resolution


First author:
M. DiDonato
Resolution: 2.30 Å
R/Rfree: 0.22/0.30
Resolution: 2.30 Å
R/Rfree: 0.22/0.30
X-ray diffraction data for the Crystal structure of 30S ribosomal protein S6 (TM0603) from Thermotoga maritima at 1.70 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.18/0.24
Resolution: 1.70 Å
R/Rfree: 0.18/0.24
X-ray diffraction data for the Crystal structure of a thioesterase-like protein (tm0581) from thermotoga maritima msb8 at 2.20 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.20 Å
R/Rfree: 0.18/0.22
Resolution: 2.20 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of S-adenosylmethionine tRNA ribosyltransferase (TM0574) from Thermotoga maritima at 2.00 A resolution


First author:
I. Mathews
Resolution: 2.00 Å
R/Rfree: 0.18/0.21
Resolution: 2.00 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of Cell division protein ftsY (TM0570) from Thermotoga maritima at 1.60 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.60 Å
R/Rfree: 0.21/0.25
Resolution: 1.60 Å
R/Rfree: 0.21/0.25
X-ray diffraction data for the Crystal structure of Predicted metal-dependent phosphoesterase (PHP family) (tm0559) from THERMOTOGA MARITIMA at 2.40 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.40 Å
R/Rfree: 0.17/0.22
Resolution: 2.40 Å
R/Rfree: 0.17/0.22
X-ray diffraction data for the Crystal structure of 3-isopropylmalate dehydrogenase (TM0556) from Thermotoga maritima at 1.90 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.19/0.22
Resolution: 1.90 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of ABC transporter ATP-binding protein (TM0544) from Thermotoga maritima at 2.10 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.10 Å
R/Rfree: 0.17/0.22
Resolution: 2.10 Å
R/Rfree: 0.17/0.22
X-ray diffraction data for the Crystal structure of NAD-dependent malic enzyme (TM0542) from Thermotoga maritima at 2.61 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.61 Å
R/Rfree: 0.19/0.24
Resolution: 2.61 Å
R/Rfree: 0.19/0.24
X-ray diffraction data for the Crystal structure of Uracil-DNA glycosylase (TM0511) from Thermotoga maritima at 1.90 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.18/0.21
Resolution: 1.90 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of Tryptophanyl-tRNA synthetase (EC 6.1.1.2) (Tryptophan-tRNA ligase)(TrpRS) (tm0492) from THERMOTOGA MARITIMA at 2.50 A resolution


First author:
G.W. Han
Resolution: 2.50 Å
R/Rfree: 0.19/0.26
Resolution: 2.50 Å
R/Rfree: 0.19/0.26
X-ray diffraction data for the Crystal structure of N5-glutamine methyltransferase, HemK(EC 2.1.1.-) (TM0488) from Thermotoga maritima at 2.80 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.80 Å
R/Rfree: 0.21/0.27
Resolution: 2.80 Å
R/Rfree: 0.21/0.27
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE THIAMINE BIOSYNTHESIS/SALVAGE PROTEIN (TM0486) FROM THERMOTOGA MARITIMA AT 1.80 A RESOLUTION


First author:
Z. Dermoun
Resolution: 1.80 Å
R/Rfree: 0.14/0.19
Resolution: 1.80 Å
R/Rfree: 0.14/0.19
X-ray diffraction data for the Crystal structure of the catalytic subunit of a phosphoribosylaminoimidazole mutase (tm0446) from thermotoga maritima at 1.77 A resolution


First author:
R. Schwarzenbacher
Resolution: 1.77 Å
R/Rfree: 0.15/0.18
Resolution: 1.77 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of Gluconate 5-dehydrogenase (TM0441) from Thermotoga maritima at 2.07 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.07 Å
R/Rfree: 0.15/0.18
Resolution: 2.07 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of Alcohol dehydrogenase (TM0436) from Thermotoga maritima at 2.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.15/0.19
Resolution: 2.00 Å
R/Rfree: 0.15/0.19
X-ray diffraction data for the Crystal structure of PfkB Carbohydrate kinase (TM0415) from Thermotoga maritima at 1.91 A resolution

First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.91 Å
R/Rfree: 0.17/0.22
Resolution: 1.91 Å
R/Rfree: 0.17/0.22
X-ray diffraction data for the Crystal structure of glycerol-3-phosphate dehydrogenase (TM0378) from THERMOTOGA MARITIMA at 2.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.17/0.21
Resolution: 2.00 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of Phospho-2-dehydro-3-deoxyheptonate aldolase (DAHP synthase) (TM0343) from Thermotoga Maritima at 1.92 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.92 Å
R/Rfree: 0.17/0.22
Resolution: 1.92 Å
R/Rfree: 0.17/0.22
X-ray diffraction data for the Crystal structure of Orotidine 5'-phosphate decarboxylase (TM0332) from Thermotoga maritima at 2.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.18/0.22
Resolution: 2.00 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of Oxidoreductase (TM0312) from Thermotoga maritima at 2.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.50 Å
R/Rfree: 0.19/0.24
Resolution: 2.50 Å
R/Rfree: 0.19/0.24