415 results
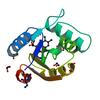
X-ray diffraction data for the Crystal structure of Flavoprotein in Complex with FMN (YP_193882.1) from Lactobacillus acidophilus NCFM at 1.20 A resolution

First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.20 Å
R/Rfree: 0.14/0.16
Resolution: 1.20 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the Crystal structure of a putative superoxide reductase (TM0658) from THERMOTOGA MARITIMA at 1.10 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.10 Å
R/Rfree: 0.12/0.15
Resolution: 1.10 Å
R/Rfree: 0.12/0.15
X-ray diffraction data for the Crystal structure of an a putative hydrolase of the isochorismatase family (CV_1320) from Chromobacterium violaceum ATCC 12472 at 1.06 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.06 Å
R/Rfree: 0.13/0.15
Resolution: 1.06 Å
R/Rfree: 0.13/0.15
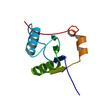
X-ray diffraction data for the Room temperature crystal structure of a RNA binding motif protein 39 (Rbm39) from Mus musculus at 1.11 A resolution

First author:
Partnership for T-Cell Biology (TCELL) JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.11 Å
R/Rfree: 0.11/0.13
Resolution: 1.11 Å
R/Rfree: 0.11/0.13
X-ray diffraction data for the Crystal structure of a putative flavin reductase (ycdh, hs_1225) from haemophilus somnus 129pt at 1.06 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.06 Å
R/Rfree: 0.14/0.17
Resolution: 1.06 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of a putative acetyltransferase (DR_1678) from Deinococcus radiodurans R1 at 1.19 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.19 Å
R/Rfree: 0.14/0.15
Resolution: 1.19 Å
R/Rfree: 0.14/0.15
X-ray diffraction data for the Crystal structure of a putative endopeptidase (ava_3396) from anabaena variabilis atcc 29413 at 1.05 A resolution


First author:
Q. Xu
Resolution: 1.05 Å
R/Rfree: 0.13/0.15
Resolution: 1.05 Å
R/Rfree: 0.13/0.15
X-ray diffraction data for the Crystal structure of a Histidine triad protein (Maqu_1709) from Marinobacter aquaeolei VT8 at 1.20 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.20 Å
R/Rfree: 0.13/0.15
Resolution: 1.20 Å
R/Rfree: 0.13/0.15
X-ray diffraction data for the Crystal structure of a putative s-adenosyl-l-methionine-dependent methyltransferase (mmp1179) from methanococcus maripaludis at 1.15 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.15 Å
R/Rfree: 0.12/0.14
Resolution: 1.15 Å
R/Rfree: 0.12/0.14
X-ray diffraction data for the Crystal structure of a DUF1989 family protein (TM1040_0329) from SILICIBACTER SP. TM1040 at 1.11 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.11 Å
R/Rfree: 0.13/0.15
Resolution: 1.11 Å
R/Rfree: 0.13/0.15
X-ray diffraction data for the Crystal structure of a ftsz-like protein of unknown function (npun_r1471) from nostoc punctiforme pcc 73102 at 1.22 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.22 Å
R/Rfree: 0.16/0.18
Resolution: 1.22 Å
R/Rfree: 0.16/0.18
X-ray diffraction data for the Crystal structure of a RNA binding motif protein 39 (RBM39) from Mus musculuS at 0.95 A resolution


First author:
Partnership for T-Cell Biology (TCELL) Joint Center for Structural Genomics (JCSG)
Resolution: 0.95 Å
R/Rfree: 0.12/0.13
Resolution: 0.95 Å
R/Rfree: 0.12/0.13
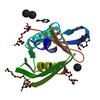
X-ray diffraction data for the Crystal structure of a duf4847 family protein (BVU_2626) from Bacteroides vulgatus ATCC 8482 at 1.00 A resolution

First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.00 Å
R/Rfree: 0.12/0.14
Resolution: 1.00 Å
R/Rfree: 0.12/0.14
X-ray diffraction data for the Crystal structure of a putative hydrolase (lpg1103) from Legionella pneumophila subsp. pneumophila str. Philadelphia 1 at 1.15 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.15 Å
R/Rfree: 0.14/0.16
Resolution: 1.15 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the Crystal structure of a putative nucleotidyltransferase (tm1012) from Thermotoga maritima at 1.35 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.20 Å
R/Rfree: 0.14/0.17
Resolution: 1.20 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of an ankyrin repeat domain (ABAYE2397) from Acinetobacter baumannii AYE at 1.00 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.00 Å
R/Rfree: 0.14/0.17
Resolution: 1.00 Å
R/Rfree: 0.14/0.17
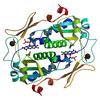
X-ray diffraction data for the Crystal structure of a putative fmn-dependent nitroreductase (ct0345) from chlorobium tepidum tls at 1.15 A resolution

First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.15 Å
R/Rfree: 0.15/0.17
Resolution: 1.15 Å
R/Rfree: 0.15/0.17
X-ray diffraction data for the Crystal structure of protein of unknown function (DUF1255) (AFE_2634) from ACIDITHIOBACILLUS FERROOXIDANS NCIB8455 at 0.97 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 0.97 Å
R/Rfree: 0.12/0.14
Resolution: 0.97 Å
R/Rfree: 0.12/0.14
X-ray diffraction data for the Crystal structure of Na(+)-translocating NADH-quinone reductase subunit C (YP_001302508.1) from Parabacteroides distasonis ATCC 8503 at 1.10 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.10 Å
R/Rfree: 0.12/0.14
Resolution: 1.10 Å
R/Rfree: 0.12/0.14
X-ray diffraction data for the Crystal structure of a N-acetylmuramoyl-L-alanine amidase (BACUNI_02947) from Bacteroides uniformis ATCC 8492 at 1.15 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.15 Å
R/Rfree: 0.13/0.16
Resolution: 1.15 Å
R/Rfree: 0.13/0.16
X-ray diffraction data for the Crystal structure of a putative polyketide cyclase (pp0894) from pseudomonas putida kt2440 at 1.24 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.24 Å
R/Rfree: 0.12/0.15
Resolution: 1.24 Å
R/Rfree: 0.12/0.15
X-ray diffraction data for the Crystal structure of a N-acetylmuramoyl-L-alanine amidase (BACUNI_02947) from Bacteroides uniformis ATCC 8492 at 1.07 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.07 Å
R/Rfree: 0.11/0.14
Resolution: 1.07 Å
R/Rfree: 0.11/0.14
X-ray diffraction data for the Crystal structure of a putative isochorismatase (Bxe_A0706) from BURKHOLDERIA XENOVORANS LB400 at 1.22 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.22 Å
R/Rfree: 0.14/0.17
Resolution: 1.22 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of S-adenosylmethionine decarboxylase proenzyme (TM0655) from THERMOTOGA MARITIMA at 1.2 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.20 Å
Resolution: 1.20 Å
X-ray diffraction data for the Co-crystal Structure of 3-hydroxydecanoyl-(acyl carrier protein) Dehydratase from Yersinia pestis with 5-Benzoylpentanoic Acid

First author:
N. Maltseva
Gene name: fabA
Resolution: 1.18 Å
R/Rfree: 0.15/0.17
Gene name: fabA
Resolution: 1.18 Å
R/Rfree: 0.15/0.17
X-ray diffraction data for the Crystal Structure of a Phosphocarrier Protein HPr from Bacillus anthracis str. Ames

First author:
J.S. Brunzelle
Gene name: ptsH-2
Resolution: 1.15 Å
R/Rfree: 0.16/0.18
Gene name: ptsH-2
Resolution: 1.15 Å
R/Rfree: 0.16/0.18
X-ray diffraction data for the 0.95A Resolution Structure of a Histidine Triad Protein from Clostridium difficile

First author:
S.M. Anderson
Resolution: 0.95 Å
R/Rfree: 0.14/0.15
Resolution: 0.95 Å
R/Rfree: 0.14/0.15
X-ray diffraction data for the Structure of phosphotransferase enzyme II, A component from Yersinia pestis CO92 at 1.2 A resolution

First author:
E.V. Filippova
Resolution: 1.20 Å
R/Rfree: 0.15/0.18
Resolution: 1.20 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the 1.05 Angstrom crystal structure of an amino acid ABC transporter substrate-binding protein AbpA from Streptococcus pneumoniae Canada MDR_19A bound to L-arginine

First author:
P.J. Stogios
Resolution: 1.05 Å
R/Rfree: 0.17/0.18
Resolution: 1.05 Å
R/Rfree: 0.17/0.18
X-ray diffraction data for the 1.2 Angstrom Crystal Structure of the Glutaredoxin 2 (grxB) from Salmonella typhimurium in complex with Glutathione

First author:
G. Minasov
Gene name: grxB
Resolution: 1.20 Å
R/Rfree: 0.12/0.14
Gene name: grxB
Resolution: 1.20 Å
R/Rfree: 0.12/0.14
X-ray diffraction data for the Crystal structure of PriA from Actinomyces urogenitalis


First author:
K. MICHALSKA
Resolution: 1.05 Å
R/Rfree: 0.12/0.14
Resolution: 1.05 Å
R/Rfree: 0.12/0.14
X-ray diffraction data for the 1.18 Angstrom resolution crystal structure of uncharacterized protein lmo1340 from Listeria monocytogenes EGD-e

First author:
A.S. Halavaty
Gene name: -
Resolution: 1.18 Å
R/Rfree: 0.15/0.18
Gene name: -
Resolution: 1.18 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the 1.1 Angstrom Crystal Structure of Hypothetical Protein BA_2335 from Bacillus anthracis

First author:
G. Minasov
Resolution: 1.10 Å
R/Rfree: 0.15/0.18
Resolution: 1.10 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of VC2308 protein

First author:
E. Niedzialkowska
Resolution: 1.16 Å
R/Rfree: 0.14/0.17
Resolution: 1.16 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of a putative extracellular heme-binding protein (DESPIG_02683) from Desulfovibrio piger ATCC 29098 at 1.24 A resolution

First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.24 Å
R/Rfree: 0.13/0.15
Resolution: 1.24 Å
R/Rfree: 0.13/0.15
X-ray diffraction data for the Crystal structure of SusD superfamily protein (YP_001299712.1) from Bacteroides vulgatus ATCC 8482 at 1.13 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.13 Å
R/Rfree: 0.12/0.14
Resolution: 1.13 Å
R/Rfree: 0.12/0.14
X-ray diffraction data for the PKD domain of M14-like peptidase from Thermoplasmatales archaeon SCGC AB-540-F20


First author:
K. Michalska
Resolution: 1.23 Å
R/Rfree: 0.14/0.17
Resolution: 1.23 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Structure of the YrdA ferripyochelin binding protein from Salmonella enterica

First author:
S.M. Anderson
Gene name: yrdA
Resolution: 1.20 Å
R/Rfree: 0.14/0.16
Gene name: yrdA
Resolution: 1.20 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the New Delhi Metallo-beta-Lactamase-1 1.05 A structure Complexed with Hydrolyzed Ampicillin


First author:
Y. Kim
Resolution: 1.05 Å
R/Rfree: 0.13/0.16
Resolution: 1.05 Å
R/Rfree: 0.13/0.16
X-ray diffraction data for the Crystal structure of hypothetical protein with ketosteroid isomerase-like protein fold from Catenulispora acidiphila DSM 44928


First author:
E.V. Filippova
Resolution: 1.15 Å
R/Rfree: 0.13/0.14
Resolution: 1.15 Å
R/Rfree: 0.13/0.14
X-ray diffraction data for the Crystal structure of TrpF from Jonesia denitrificans


First author:
K. Michalska
Resolution: 1.09 Å
R/Rfree: 0.13/0.14
Resolution: 1.09 Å
R/Rfree: 0.13/0.14
X-ray diffraction data for the Fatty acid ABC transporter substrate-binding protein from Thermomonospora curvata


First author:
J. Osipiuk
Resolution: 1.15 Å
R/Rfree: 0.12/0.15
Resolution: 1.15 Å
R/Rfree: 0.12/0.15
X-ray diffraction data for the 1.03 Angstrom Crystal Structure of Q236A Mutant Type I Dehydroquinate Dehydratase (aroD) from Salmonella typhimurium

First author:
S.H. Light
Gene name: aroD
Resolution: 1.03 Å
R/Rfree: 0.14/0.16
Gene name: aroD
Resolution: 1.03 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the Crystal Structure of ThiJ/PfpI Domain Protein from Brachyspira murdochii


First author:
Y. Kim
Resolution: 1.12 Å
R/Rfree: 0.12/0.13
Resolution: 1.12 Å
R/Rfree: 0.12/0.13
X-ray diffraction data for the Crystal structure of a putative ornithine aminotransferase from Toxoplasma gondii ME49 in complex with pyrodoxal-5'-phosphate

First author:
E.V. Filippova
Resolution: 1.20 Å
R/Rfree: 0.13/0.16
Resolution: 1.20 Å
R/Rfree: 0.13/0.16
X-ray diffraction data for the Crystal structure of transcriptional regulator VanUg, Form II

First author:
P.J. Stogios
Gene name: vanUG
Resolution: 1.12 Å
R/Rfree: 0.14/0.16
Gene name: vanUG
Resolution: 1.12 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the 1.0 Angstrom resolution crystal structure of the branched-chain amino acid transporter substrate binding protein LivJ from Streptococcus pneumoniae str. Canada MDR_19A in complex with Isoleucine

First author:
A.S. Halavaty
Resolution: 1.00 Å
R/Rfree: 0.13/0.15
Resolution: 1.00 Å
R/Rfree: 0.13/0.15
X-ray diffraction data for the Crystal structure of iron uptake ABC transporter substrate-binding protein PiuA from Streptococcus pneumoniae Canada MDR_19A

First author:
P.J. Stogios
Resolution: 1.13 Å
R/Rfree: 0.14/0.17
Resolution: 1.13 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the 1.02 Angstrom resolution crystal structure of 3-phosphoshikimate 1-carboxyvinyltransferase from Vibrio cholerae in complex with shikimate-3-phosphate (partially photolyzed) and glyphosate

First author:
G. Minasov
Resolution: 1.02 Å
R/Rfree: 0.11/0.14
Resolution: 1.02 Å
R/Rfree: 0.11/0.14
X-ray diffraction data for the 1.1 Angstrom Crystal Structure of Putative Modulator of Drug Activity (MdaB) from Yersinia pestis CO92.

First author:
G. Minasov
Gene name: mdaB
Resolution: 1.10 Å
R/Rfree: 0.10/0.12
Gene name: mdaB
Resolution: 1.10 Å
R/Rfree: 0.10/0.12
X-ray diffraction data for the Structure of a NADH-dependent enoyl-ACP reductase from Acinetobacter baumannii in complex with NAD


First author:
J. Abendroth
Resolution: 1.20 Å
R/Rfree: 0.13/0.14
Resolution: 1.20 Å
R/Rfree: 0.13/0.14
X-ray diffraction data for the X-ray crystal structure of thioredoxin from Mycobacterium avium


First author:
C.M. Lukacs
Resolution: 1.20 Å
R/Rfree: 0.16/0.18
Resolution: 1.20 Å
R/Rfree: 0.16/0.18
X-ray diffraction data for the Crystal Structure of a Beta-1,3-glucanase from Mycobacterium marinum

First author:
D.M. Dranow
Resolution: 1.20 Å
R/Rfree: 0.12/0.14
Resolution: 1.20 Å
R/Rfree: 0.12/0.14
X-ray diffraction data for the Crystal structure of a putative endoribonuclease L-PSP from Entamoeba histolytica, rhomobohedral form


First author:
A. Gardberg
Resolution: 1.20 Å
R/Rfree: 0.11/0.13
Resolution: 1.20 Å
R/Rfree: 0.11/0.13
X-ray diffraction data for the New ligand for thaumatin discovered using acoustic high throughput screening


First author:
E. Teplitsky
Resolution: 1.20 Å
R/Rfree: 0.15/0.16
Resolution: 1.20 Å
R/Rfree: 0.15/0.16
X-ray diffraction data for the Crystal structure of a GNAT superfamily phosphinothricin acetyltransferase (Pat) from Sinorhizobium meliloti 1021


First author:
K.A. Majorek
Resolution: 1.15 Å
R/Rfree: 0.12/0.15
Resolution: 1.15 Å
R/Rfree: 0.12/0.15
X-ray diffraction data for the Crystal structure of a GNAT superfamily phosphinothricin acetyltransferase (Pat) from Sinorhizobium meliloti in complex with AcCoA


First author:
K.A. Majorek
Resolution: 1.15 Å
R/Rfree: 0.12/0.14
Resolution: 1.15 Å
R/Rfree: 0.12/0.14
X-ray diffraction data for the Crystal structure of a DUF4467 family protein (SAV0303) from Staphylococcus aureus subsp. aureus Mu50 at 1.35 A resolution

First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.20 Å
R/Rfree: 0.14/0.17
Resolution: 1.20 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of a leucine-rich repeat protein (BACOVA_04585) from Bacteroides ovatus ATCC 8483 at 1.19 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.19 Å
R/Rfree: 0.13/0.16
Resolution: 1.19 Å
R/Rfree: 0.13/0.16
X-ray diffraction data for the Crystal structure of a DJ-1 (PARK7) from Homo sapiens at 1.23 A resolution


First author:
Partnership for Nuclear Receptor Signaling Code Biology (NHRs) Joint Center for Structural Genomics (JCSG)
Resolution: 1.23 Å
R/Rfree: 0.12/0.13
Resolution: 1.23 Å
R/Rfree: 0.12/0.13
X-ray diffraction data for the Crystal structure of a RNA binding domain of poly-U binding splicing factor 60KDa (PUF60) from Homo sapiens at 1.23 A resolution


First author:
Partnership for T-Cell Biology JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.23 Å
R/Rfree: 0.13/0.16
Resolution: 1.23 Å
R/Rfree: 0.13/0.16
X-ray diffraction data for the Crystal structure of transaldolase (YP_208650.1) from Neisseria gonorrhoeae FA 1090 at 1.14 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.14 Å
R/Rfree: 0.13/0.16
Resolution: 1.14 Å
R/Rfree: 0.13/0.16
X-ray diffraction data for the Crystal structure of a duf1831 family protein (lp2179) from lactobacillus plantarum at 1.20 A resolution

First author:
C. Bakolitsa
Resolution: 1.20 Å
R/Rfree: 0.12/0.15
Resolution: 1.20 Å
R/Rfree: 0.12/0.15
X-ray diffraction data for the Crystal structure of an ntf2-like protein with a cystatin-like fold (saro_3722) from novosphingobium aromaticivorans dsm at 1.16 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.16 Å
R/Rfree: 0.14/0.17
Resolution: 1.16 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of Saro_0823 (YP_496102.1) a protein of unknown function from NOVOSPHINGOBIUM AROMATICIVORANS DSM 12444 at 1.22 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.22 Å
R/Rfree: 0.12/0.15
Resolution: 1.22 Å
R/Rfree: 0.12/0.15
X-ray diffraction data for the Crystal structure of a BLIP-like protein (BF1215) from Bacteroides fragilis NCTC 9343 at 1.20 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.20 Å
R/Rfree: 0.15/0.17
Resolution: 1.20 Å
R/Rfree: 0.15/0.17
X-ray diffraction data for the Crystal structure of a Zn-dependent exopeptidase (BDI_3547) from Parabacteroides distasonis ATCC 8503 at 1.06 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.06 Å
R/Rfree: 0.11/0.13
Resolution: 1.06 Å
R/Rfree: 0.11/0.13
X-ray diffraction data for the Crystal structure of a putative signal transduction protein (eubrec_0645) from eubacterium rectale atcc 33656 at 1.20 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.20 Å
R/Rfree: 0.12/0.14
Resolution: 1.20 Å
R/Rfree: 0.12/0.14
X-ray diffraction data for the Crystal structure of a putative glucosamine-fructose-6-phosphate aminotransferase (stm4540.s) from salmonella typhimurium lt2 at 1.35 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.23 Å
R/Rfree: 0.11/0.14
Resolution: 1.23 Å
R/Rfree: 0.11/0.14
X-ray diffraction data for the Crystal structure of Putative reductase (NP_038806.2) from MUS MUSCULUS at 1.18 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.18 Å
R/Rfree: 0.13/0.16
Resolution: 1.18 Å
R/Rfree: 0.13/0.16
X-ray diffraction data for the Crystal structure of a low occupancy fragment candidate (5-[(4-Isopropylphenyl)amino]-6-methyl-1,2,4-triazin-3(2H)-one) bound in the ubiquitin binding pocket of the HDAC6 zinc-finger domain

First author:
R.J. Harding
Resolution: 1.07 Å
R/Rfree: 0.14/0.15
Resolution: 1.07 Å
R/Rfree: 0.14/0.15
X-ray diffraction data for the Crystal structure of a low occupancy fragment candidate (N-(4-Methyl-1,3-thiazol-2-yl)propanamide) bound adjacent to the ubiquitin binding pocket of the HDAC6 zinc-finger domain

First author:
R.J. Harding
Resolution: 1.05 Å
R/Rfree: 0.13/0.15
Resolution: 1.05 Å
R/Rfree: 0.13/0.15
X-ray diffraction data for the Crystal structure of a NAD-dependent Aldehyde dehydrogenase from Burkholderia multivorans in covalent complex with NAD


First author:
J. Abendroth
Resolution: 1.20 Å
R/Rfree: 0.11/0.13
Resolution: 1.20 Å
R/Rfree: 0.11/0.13
X-ray diffraction data for the Crystal structure of a beta-lactamase from Burkholderia ambifaria


First author:
S.J. Mayclin
Resolution: 1.10 Å
R/Rfree: 0.14/0.17
Resolution: 1.10 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of cyclohexanone monooxygenase from Thermocrispum municipale in the oxidised state with a bound nicotinamide.


First author:
J.R. Gomez-Castellanos
Resolution: 1.22 Å
R/Rfree: 0.13/0.17
Resolution: 1.22 Å
R/Rfree: 0.13/0.17
X-ray diffraction data for the Crystal structure of Lupinus luteus S-adenosyl-L-homocysteine hydrolase in complex with adenosine


First author:
K. Brzezinski
Resolution: 1.17 Å
R/Rfree: 0.13/0.16
Resolution: 1.17 Å
R/Rfree: 0.13/0.16
X-ray diffraction data for the Crystal structure of 5'-methylthioadenosine/S-adenosylhomocysteine nucleosidase in complex with adenine from Vibrio fischeri ES114


X-ray diffraction data for the MdbA protein, a thiol-disulfide oxidoreductase from Corynebacterium matruchotii

First author:
J. Osipiuk
Resolution: 1.20 Å
R/Rfree: 0.12/0.15
Resolution: 1.20 Å
R/Rfree: 0.12/0.15
X-ray diffraction data for the Crystal structure of a thioredoxin domain 2 from Brucella melitensis at 1.15 Angstrom resolution


First author:
S.J. Mayclin
Resolution: 1.15 Å
R/Rfree: 0.14/0.16
Resolution: 1.15 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the Crystal structure of a methyltransferase


First author:
C. Dong
Resolution: 1.20 Å
R/Rfree: 0.15/0.18
Resolution: 1.20 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the 1.05 Angstrom Resolution Crystal Structure of UDP-N-acetylglucosamine 1-carboxyvinyltransferase from Acinetobacter baumannii in Covalently Bound Complex with (2R)-2-(phosphonooxy)propanoic Acid.


X-ray diffraction data for the Catalytic domain of LPMO Lmo2467 from Listeria monocytogenes


X-ray diffraction data for the 1.0 Angstrom Crystal Structure of pre-Peptidase C-terminal Domain of Collagenase from Bacillus anthracis.


X-ray diffraction data for the Ultra-high resolution structure of d(CGCGCG)2 Z-DNA


First author:
K. Brzezinski
Resolution: 0.55 Å
R/Rfree: 0.08/0.09
Resolution: 0.55 Å
R/Rfree: 0.08/0.09
X-ray diffraction data for the Dimeric structure of the N-terminal domain of PriB protein from Thermoanaerobacter tencongensis solved ab initio


First author:
D. Liebschner
Resolution: 1.09 Å
Resolution: 1.09 Å
X-ray diffraction data for the Beta-lactamase penicillinase from Bacillus megaterium


X-ray diffraction data for the Crystal Structure of Ribose-5-phosphate Isomerase B from Mycoplasma genitalium with bound Ribulose-5-phosphate


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.10 Å
R/Rfree: 0.13/0.15
Resolution: 1.10 Å
R/Rfree: 0.13/0.15
X-ray diffraction data for the Structure of the engineered metalloesterase MID1sc9


First author:
S. Studer
Resolution: 1.13 Å
R/Rfree: 0.15/0.19
Resolution: 1.13 Å
R/Rfree: 0.15/0.19
X-ray diffraction data for the beta-lactamase SHV-11 from Klebsiella pneumoniae strain NTUH-K2044


First author:
J. Osipiuk
Gene name: blaSHV-11
Resolution: 1.17 Å
R/Rfree: 0.12/0.13
Gene name: blaSHV-11
Resolution: 1.17 Å
R/Rfree: 0.12/0.13
X-ray diffraction data for the Crystal Structure of chromodomain of CBX8 in complex with inhibitor UNC3866

First author:
Y. Liu
Resolution: 1.18 Å
R/Rfree: 0.16/0.19
Resolution: 1.18 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the 1.16 Angstrom Resolution Crystal Structure of Acyl Carrier Protein Domain (residues 1-100) of Polyketide Synthase Pks13 from Mycobacterium tuberculosis


X-ray diffraction data for the 1.2 Angstrom Resolution Crystal Structure of Nucleoside Triphosphatase NudI from Klebsiella pneumoniae in Complex with HEPES


X-ray diffraction data for the Crystal structure of the S65T/F99S/M153T/V163A variant of GFP at 0.85 A


First author:
Y. Tai
Resolution: 0.85 Å
R/Rfree: 0.09/0.11
Resolution: 0.85 Å
R/Rfree: 0.09/0.11
X-ray diffraction data for the Crystal structure of the F99S/M153T/V163A/T203I variant of GFP at 0.94 A


First author:
H. Eki
Resolution: 0.94 Å
R/Rfree: 0.11/0.13
Resolution: 0.94 Å
R/Rfree: 0.11/0.13
X-ray diffraction data for the Crystal structure of the F99S/M153T/V163A/E222Q variant of GFP at 0.78 A
First author:
K. Takaba
Resolution: 0.78 Å
R/Rfree: 0.11/0.13
Resolution: 0.78 Å
R/Rfree: 0.11/0.13
X-ray diffraction data for the FimA wt from S. flexneri


First author:
D. Zyla
Resolution: 0.89 Å
R/Rfree: 0.11/0.13
Resolution: 0.89 Å
R/Rfree: 0.11/0.13
X-ray diffraction data for the Crystal structure of HLA-B*2709 complexed with the glucagon receptor (GR) peptide (residues 412-420)


First author:
B. Loll
Resolution: 1.20 Å
R/Rfree: 0.13/0.15
Resolution: 1.20 Å
R/Rfree: 0.13/0.15
X-ray diffraction data for the Crystal Structure of a Probable short-chain type dehydrogenase/reductase (Rv1144) from Mycobacterium tuberculosis with bound NAD


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.20 Å
R/Rfree: 0.12/0.14
Resolution: 1.20 Å
R/Rfree: 0.12/0.14
X-ray diffraction data for the HhaI endonuclease in Complex with DNA at 1 Angstrom Resolution


First author:
J.R. Horton
Resolution: 1.00 Å
R/Rfree: 0.16/0.17
Resolution: 1.00 Å
R/Rfree: 0.16/0.17
X-ray diffraction data for the HhaI endonuclease in Complex With DNA in space group P21 (pH 4.2)


First author:
J.R. Horton
Resolution: 1.16 Å
R/Rfree: 0.14/0.16
Resolution: 1.16 Å
R/Rfree: 0.14/0.16