6850 results
X-ray diffraction data for the Crystal structure of Putative carboxypeptidase A (YP_562911.1) from SHEWANELLA DENITRIFICANS OS-217 at 2.39 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.39 Å
R/Rfree: 0.18/0.23
Resolution: 2.39 Å
R/Rfree: 0.18/0.23
X-ray diffraction data for the Crystal structure of putative carboxypeptidase (YP_103406.1) from BURKHOLDERIA MALLEI ATCC 23344 at 2.49 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.49 Å
R/Rfree: 0.16/0.19
Resolution: 2.49 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of a putative zinc peptidase (NP_812461.1) from Bacteroides thetaiotaomicron VPI-5482 at 2.31 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.31 Å
R/Rfree: 0.19/0.25
Resolution: 2.31 Å
R/Rfree: 0.19/0.25
X-ray diffraction data for the Crystal structure of putative peptidase (NP_348812.1) from CLOSTRIDIUM ACETOBUTYLICUM at 2.37 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.37 Å
R/Rfree: 0.17/0.21
Resolution: 2.37 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of a succinylglutamate desuccinylase (TM1040_2694) from SILICIBACTER SP. TM1040 at 2.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.16/0.20
Resolution: 2.00 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the Crystal structure of an aspartoacylase family protein (mlr6093) from mesorhizobium loti maff303099 at 2.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.21/0.25
Resolution: 2.00 Å
R/Rfree: 0.21/0.25
X-ray diffraction data for the Crystal Structure of the SRAP Domain of Human HMCES Protein


First author:
L. Halabelian
Resolution: 1.50 Å
R/Rfree: 0.18/0.21
Resolution: 1.50 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the The tetramer structure of the nervy homolgy two (NHR2) domain of AML1-ETO is critical for AML1-ETO'S activity


First author:
Y. Liu
Resolution: 2.00 Å
R/Rfree: 0.21/0.27
Resolution: 2.00 Å
R/Rfree: 0.21/0.27
X-ray diffraction data for the Crystal structure of carbon-nitrogen hydrolase from Helicobacter pylori G27


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 2.10 Å
R/Rfree: 0.17/0.21
Resolution: 2.10 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of a low occupancy fragment candidate (5-[(4-Isopropylphenyl)amino]-6-methyl-1,2,4-triazin-3(2H)-one) bound in the ubiquitin binding pocket of the HDAC6 zinc-finger domain

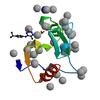
First author:
R.J. Harding
Resolution: 1.07 Å
R/Rfree: 0.14/0.15
Resolution: 1.07 Å
R/Rfree: 0.14/0.15
X-ray diffraction data for the Crystal structure of a low occupancy fragment candidate (N-(4-Methyl-1,3-thiazol-2-yl)propanamide) bound adjacent to the ubiquitin binding pocket of the HDAC6 zinc-finger domain

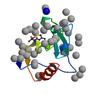
First author:
R.J. Harding
Resolution: 1.05 Å
R/Rfree: 0.13/0.15
Resolution: 1.05 Å
R/Rfree: 0.13/0.15
X-ray diffraction data for the Crystal structure of fragment (3-[6-Oxo-3-(3-pyridinyl)-1(6H)-pyridazinyl]propanoic acid) bound in the ubiquitin binding pocket of the HDAC6 zinc-finger domain

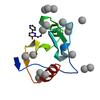
First author:
R.J. Harding
Resolution: 1.70 Å
R/Rfree: 0.15/0.18
Resolution: 1.70 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of fragment (3-(5-Chloro-1,3-benzothiazol-2-yl)propanoic acid) bound in the ubiquitin binding pocket of the HDAC6 zinc-finger domain

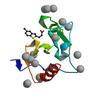
First author:
R.J. Harding
Resolution: 1.60 Å
R/Rfree: 0.16/0.19
Resolution: 1.60 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of fragment 3-(1,3-Benzothiazol-2-yl)propanoic acid bound in the ubiquitin binding pocket of the HDAC6 zinc-finger domain

First author:
R.J. Harding
Resolution: 1.80 Å
R/Rfree: 0.15/0.18
Resolution: 1.80 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of fragment 3-(1-Methyl-2-oxo-1,2-dihydroquinoxalin-3-yl)propionic acid bound in the ubiquitin binding pocket of the HDAC6 zinc-finger domain

First author:
R.J. Harding
Resolution: 1.55 Å
R/Rfree: 0.15/0.17
Resolution: 1.55 Å
R/Rfree: 0.15/0.17