6726 results
X-ray diffraction data for the CRYSTAL STRUCTURE OF A NTF2-like protein (BTH_I0051) FROM BURKHOLDERIA THAILANDENSIS E264 AT 1.60 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.60 Å
R/Rfree: 0.17/0.20
Resolution: 1.60 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of NTF2-superfamily protein with unknown function (NP_977240.1) from BACILLUS CEREUS ATCC 10987 at 1.25 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.25 Å
R/Rfree: 0.13/0.17
Resolution: 1.25 Å
R/Rfree: 0.13/0.17
X-ray diffraction data for the Crystal structure of Putative PLP-dependent aminotransferase (NP_978343.1) from Bacillus cereus ATCC 10987 at 2.19 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.19 Å
R/Rfree: 0.18/0.22
Resolution: 2.19 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of a putative nitroreductase (ava_2154) from anabaena variabilis atcc 29413 at 1.80 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.80 Å
R/Rfree: 0.14/0.16
Resolution: 1.80 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the Crystal structure of an ntf2-like protein (ava_4193) from anabaena variabilis atcc 29413 at 1.90 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.18/0.22
Resolution: 1.90 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of Protein of unknown function (DUF861) with a RmlC-like cupin fold (17741406) from AGROBACTERIUM TUMEFACIENS str. C58 (Dupont) at 1.64 A resolution

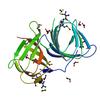
First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.64 Å
R/Rfree: 0.14/0.16
Resolution: 1.64 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the Crystal structure of a snoal-like polyketide cyclase (atu3018) from agrobacterium tumefaciens str. c58 at 2.12 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.12 Å
R/Rfree: 0.21/0.24
Resolution: 2.12 Å
R/Rfree: 0.21/0.24
X-ray diffraction data for the Crystal structure of uncharacterized protein conserved in bacteria with a cystatin-like fold (YP_168589.1) from SILICIBACTER POMEROYI DSS-3 at 2.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.18/0.19
Resolution: 2.00 Å
R/Rfree: 0.18/0.19
X-ray diffraction data for the Crystal structure of a STEROID DELTA-ISOMERASE (NP_250810.1) from PSEUDOMONAS AERUGINOSA at 2.57 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.57 Å
R/Rfree: 0.19/0.24
Resolution: 2.57 Å
R/Rfree: 0.19/0.24
X-ray diffraction data for the Crystal structure of a putative delta-5-3-ketosteroid isomerase (eca2236) from pectobacterium atrosepticum scri1043 at 1.55 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.55 Å
R/Rfree: 0.16/0.19
Resolution: 1.55 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE ANTIBIOTIC BIOSYNTHESIS MONOOXYGENASE (DR_2100) FROM DEINOCOCCUS RADIODURANS AT 1.40 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.40 Å
R/Rfree: 0.13/0.16
Resolution: 1.40 Å
R/Rfree: 0.13/0.16
X-ray diffraction data for the Crystal structure of CT0912, ORFan protein from Chlorobium tepidum with a ferredoxin-like domain repeat (NP_661805.1) from CHLOROBIUM TEPIDUM TLS at 1.80 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.80 Å
R/Rfree: 0.16/0.19
Resolution: 1.80 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of histidinol-phosphate aminotransferase (YP_297314.1) from RALSTONIA EUTROPHA JMP134 at 2.05 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.05 Å
R/Rfree: 0.19/0.24
Resolution: 2.05 Å
R/Rfree: 0.19/0.24
X-ray diffraction data for the Crystal structure of putative histidinol-phosphate aminotransferase (NP_281508.1) from Campylobacter jejuni at 2.01 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.01 Å
R/Rfree: 0.17/0.24
Resolution: 2.01 Å
R/Rfree: 0.17/0.24
X-ray diffraction data for the Crystal structure of NTF2-like protein of unknown function (YP_677363.1) from CYTOPHAGA HUTCHINSONII ATCC 33406 at 1.27 A resolution

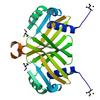
First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.27 Å
R/Rfree: 0.18/0.20
Resolution: 1.27 Å
R/Rfree: 0.18/0.20
X-ray diffraction data for the Crystal structure of NTF2-like protein of unknown function (YP_680363.1) from CYTOPHAGA HUTCHINSONII ATCC 33406 at 1.45 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.45 Å
R/Rfree: 0.17/0.18
Resolution: 1.45 Å
R/Rfree: 0.17/0.18
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PHOSPHOSERINE AMINOTRANSFERASE SERC (CHU_0995) FROM CYTOPHAGA HUTCHINSONII ATCC 33406 AT 1.75 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.75 Å
R/Rfree: 0.14/0.16
Resolution: 1.75 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the Crystal structure of putative aminotransferase (MocR family) (YP_604413.1) from DEINOCOCCUS GEOTHERMALIS DSM 11300 at 2.60 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.60 Å
R/Rfree: 0.22/0.26
Resolution: 2.60 Å
R/Rfree: 0.22/0.26
X-ray diffraction data for the Crystal structure of an aspartate transaminase (NCgl0237, Cgl0240) from CORYNEBACTERIUM GLUTAMICUM ATCC 13032 KITASATO at 1.25 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.25 Å
R/Rfree: 0.11/0.12
Resolution: 1.25 Å
R/Rfree: 0.11/0.12
X-ray diffraction data for the Crystal structure of a putative plp-dependent beta-cystathionase (aecd, dip1736) from corynebacterium diphtheriae at 1.99 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.99 Å
R/Rfree: 0.15/0.18
Resolution: 1.99 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the CRYSTAL STRUCTURE OF A FERRITIN LIKE PROTEIN (CC_0557) FROM CAULOBACTER VIBRIOIDES AT 1.95 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.95 Å
R/Rfree: 0.19/0.23
Resolution: 1.95 Å
R/Rfree: 0.19/0.23
X-ray diffraction data for the Crystal structure of nitroreductase family protein (YP_877874.1) from Clostridium novyi NT at 1.75 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.75 Å
R/Rfree: 0.17/0.22
Resolution: 1.75 Å
R/Rfree: 0.17/0.22
X-ray diffraction data for the Crystal structure of a putative ca/calmodulin-dependent kinase ii association domain (exig_1688) from exiguobacterium sibiricum 255-15 at 2.59 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.59 Å
R/Rfree: 0.20/0.24
Resolution: 2.59 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the Crystal structure of Putative aromatic amino acid beta-eliminating lyase/threonine aldolase. (YP_001813866.1) from Exiguobacterium sp. 255-15 at 2.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.14/0.19
Resolution: 2.00 Å
R/Rfree: 0.14/0.19
X-ray diffraction data for the Crystal structure of NTF2-like protein of unknown function (YP_001812677.1) from EXIGUOBACTERIUM SP. 255-15 at 1.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.50 Å
R/Rfree: 0.18/0.21
Resolution: 1.50 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of a putative tagatose-6-phosphate ketose/aldose isomerase (NT01CX_0292) from CLOSTRIDIUM NOVYI NT at 2.35 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.35 Å
R/Rfree: 0.23/0.26
Resolution: 2.35 Å
R/Rfree: 0.23/0.26
X-ray diffraction data for the Crystal structure of putative putative tagatose-6-phosphate ketose/aldose isomerase (NP_344614.1) from STREPTOCOCCUS PNEUMONIAE TIGR4 at 1.70 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.14/0.18
Resolution: 1.70 Å
R/Rfree: 0.14/0.18
X-ray diffraction data for the Crystal structure of Aminotransferase (NP_283882.1) from NEISSERIA MENINGITIDIS Z2491 at 1.91 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.91 Å
R/Rfree: 0.15/0.18
Resolution: 1.91 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of aminotransferase AspB (NP_207418.1) from HELICOBACTER PYLORI 26695 at 2.19 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.19 Å
R/Rfree: 0.16/0.22
Resolution: 2.19 Å
R/Rfree: 0.16/0.22
X-ray diffraction data for the Crystal structure of NTF-2 like protein of unknown function (YP_553245.1) from BURKHOLDERIA XENOVORANS LB400 at 1.85 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.85 Å
R/Rfree: 0.19/0.23
Resolution: 1.85 Å
R/Rfree: 0.19/0.23
X-ray diffraction data for the CRYSTAL STRUCTURE OF A NTF2-LIKE PROTEIN (BXE_B1094) FROM BURKHOLDERIA XENOVORANS LB400 AT 1.59 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.59 Å
R/Rfree: 0.18/0.21
Resolution: 1.59 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of the beta subunit of the benzoate 1,2-dioxygenase (benb, bmaa0186) from burkholderia mallei atcc 23344 at 1.90 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.19/0.22
Resolution: 1.90 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE ISOMERASE OF THE SNOAL-LIKE FAMILY (ATU_0744) FROM AGROBACTERIUM TUMEFACIENS STR. C58 AT 2.70 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.70 Å
R/Rfree: 0.23/0.26
Resolution: 2.70 Å
R/Rfree: 0.23/0.26
X-ray diffraction data for the Crystal structure of Aminotransferase (RER070207000802) from Eubacterium rectale at 1.70 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.15/0.17
Resolution: 1.70 Å
R/Rfree: 0.15/0.17
X-ray diffraction data for the Crystal structure of dimeric protein of unknown function and ferredoxin-like fold (YP_212648.1) from Bacteroides fragilis NCTC 9343 at 1.85 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.85 Å
R/Rfree: 0.17/0.21
Resolution: 1.85 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of Putative calcium/calmodulin-dependent protein kinase type II association domain (YP_315894.1) from THIOBACILLUS DENITRIFICANS ATCC 25259 at 2.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.01 Å
R/Rfree: 0.17/0.21
Resolution: 2.01 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of the beta subunit of a putative aromatic-ring-hydroxylating dioxygenase (YP_001165631.1) from NOVOSPHINGOBIUM AROMATICIVORANS DSM 12444 at 1.75 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.75 Å
R/Rfree: 0.17/0.21
Resolution: 1.75 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of valine-pyruvate aminotransferase AvtA (NP_462565.1) from Salmonella typhimurium LT2 at 1.80 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.80 Å
R/Rfree: 0.18/0.20
Resolution: 1.80 Å
R/Rfree: 0.18/0.20
X-ray diffraction data for the Crystal structure of Putative amino-acid aminotransferase (YP_265399.1) from Psychrobacter arcticum 273-4 at 2.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.50 Å
R/Rfree: 0.18/0.22
Resolution: 2.50 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of a murein peptide ligase mpl (psyc_0032) from psychrobacter arcticus 273-4 at 1.65 A resolution


First author:
D. Das
Resolution: 1.65 Å
R/Rfree: 0.15/0.19
Resolution: 1.65 Å
R/Rfree: 0.15/0.19
X-ray diffraction data for the Crystal structure of aspartate aminotransferase (E.C. 2.6.1.1) (YP_194538.1) from Lactobacillus acidophilus NCFM at 2.15 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.15 Å
R/Rfree: 0.18/0.24
Resolution: 2.15 Å
R/Rfree: 0.18/0.24
X-ray diffraction data for the Crystal structure of a putative phosphosugar isomerase (stm_0572) from salmonella typhimurium lt2 at 2.12 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.12 Å
R/Rfree: 0.17/0.21
Resolution: 2.12 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE PHOSPHOSUGAR ISOMERASE (BSU32610) FROM BACILLUS SUBTILIS AT 1.90 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.17/0.21
Resolution: 1.90 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of a putative cyclic nucleotide binding protein (spoa0323) from ruegeria pomeroyi dss-3 at 2.35 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.35 Å
R/Rfree: 0.20/0.24
Resolution: 2.35 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the Crystal structure of transcriptional regulator of Crp/Fnr family (YP_604437.1) from DEINOCOCCUS GEOTHERMALIS DSM 11300 at 1.86 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.86 Å
R/Rfree: 0.21/0.25
Resolution: 1.86 Å
R/Rfree: 0.21/0.25
X-ray diffraction data for the Crystal structure of a nitroreductase-like family protein (pnba, bh06130) from bartonella henselae str. houston-1 at 1.45 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.45 Å
R/Rfree: 0.15/0.17
Resolution: 1.45 Å
R/Rfree: 0.15/0.17
X-ray diffraction data for the Crystal structure of a putative sugar-phosphate isomerase (lmof2365_0531) from listeria monocytogenes str. 4b f2365 at 1.60 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.60 Å
R/Rfree: 0.13/0.16
Resolution: 1.60 Å
R/Rfree: 0.13/0.16
X-ray diffraction data for the Crystal structure of putative phosphosugar isomerase involved in capsule formation (YP_209877.1) from Bacteroides fragilis NCTC 9343 at 1.70 A resolution


First author:
H.J. Chiu
Resolution: 1.70 Å
R/Rfree: 0.17/0.20
Resolution: 1.70 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of a nitroreductase-like protein (smu.346) from streptococcus mutans at 1.70 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.15/0.19
Resolution: 1.70 Å
R/Rfree: 0.15/0.19
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE NAD(P)H:FMN OXIDOREDUCTASE (SE1966) FROM STAPHYLOCOCCUS EPIDERMIDIS ATCC 12228 AT 2.00 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.15/0.19
Resolution: 2.00 Å
R/Rfree: 0.15/0.19
X-ray diffraction data for the Crystal structure of a putative nad(p)h:fmn oxidoreductase (pg0310) from porphyromonas gingivalis w83 at 1.70 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.15/0.17
Resolution: 1.70 Å
R/Rfree: 0.15/0.17
X-ray diffraction data for the Crystal structure of a putative nitroreductase in complex with fmn (dde_0787) from desulfovibrio desulfuricans subsp. at 1.70 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.17/0.20
Resolution: 1.70 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE NITROREDUCTASE IN COMPLEX WITH FMN (EXIG_2970) FROM EXIGUOBACTERIUM SIBIRICUM 255-15 AT 1.85 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.85 Å
R/Rfree: 0.17/0.21
Resolution: 1.85 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of Putative NADH Oxidase (NP_348178.1) from CLOSTRIDIUM ACETOBUTYLICUM at 1.40 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.40 Å
R/Rfree: 0.13/0.17
Resolution: 1.40 Å
R/Rfree: 0.13/0.17
X-ray diffraction data for the Crystal structure of a nitroreductase family protein (cd3355) from clostridium difficile 630 at 1.58 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.58 Å
R/Rfree: 0.18/0.22
Resolution: 1.58 Å
R/Rfree: 0.18/0.22
X-ray diffraction data for the Crystal structure of a nitroreductase family protein (cd3355) from clostridium difficile 630 at 1.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.50 Å
R/Rfree: 0.14/0.17
Resolution: 1.50 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of BluB-like flavoprotein (YP_001089088.1) from CLOSTRIDIUM DIFFICILE 630 at 1.74 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.74 Å
R/Rfree: 0.16/0.20
Resolution: 1.74 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the Crystal structure of a putative nitroreductase in complex with fmn (cd3205) from clostridium difficile 630 at 1.35 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.35 Å
R/Rfree: 0.13/0.16
Resolution: 1.35 Å
R/Rfree: 0.13/0.16
X-ray diffraction data for the Crystal structure of putative oxidoreductase (YP_213212.1) from Bacteroides fragilis NCTC 9343 at 1.99 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.99 Å
R/Rfree: 0.17/0.22
Resolution: 1.99 Å
R/Rfree: 0.17/0.22
X-ray diffraction data for the Crystal structure of putative phosphosugar isomerase (YP_167080.1) from SILICIBACTER POMEROYI DSS-3 at 1.75 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.75 Å
R/Rfree: 0.16/0.20
Resolution: 1.75 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the Crystal structure of a putative phosphosugar isomerase (sden_2705) from shewanella denitrificans os217 at 2.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.17/0.20
Resolution: 2.00 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of Nitroreductase with Bound FMN (YP_211706.1) from Bacteroides fragilis NCTC 9343 at 1.70 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.15/0.18
Resolution: 1.70 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of a Nitroreductase with bound FMN (Dhaf_2018) from Desulfitobacterium hafniense DCB-2 at 2.30 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.30 Å
R/Rfree: 0.17/0.24
Resolution: 2.30 Å
R/Rfree: 0.17/0.24
X-ray diffraction data for the Human METTL3-METTL14 complex


First author:
H. ZENG
Resolution: 1.80 Å
R/Rfree: 0.20/0.22
Resolution: 1.80 Å
R/Rfree: 0.20/0.22
X-ray diffraction data for the Fragment of 7SK snRNA methylphosphate capping enzyme


First author:
H. Wu
Resolution: 2.55 Å
R/Rfree: 0.20/0.22
Resolution: 2.55 Å
R/Rfree: 0.20/0.22

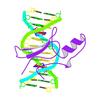
First author:
K. Liu
Resolution: 2.30 Å
R/Rfree: 0.23/0.29
Resolution: 2.30 Å
R/Rfree: 0.23/0.29
X-ray diffraction data for the Crystal structure of predicted HD superfamily hydrolase (104161995) from uncultured Thermotogales bacterium at 1.45 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.45 Å
R/Rfree: 0.18/0.19
Resolution: 1.45 Å
R/Rfree: 0.18/0.19

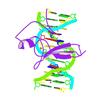
First author:
K. Liu
Resolution: 2.01 Å
R/Rfree: 0.21/0.25
Resolution: 2.01 Å
R/Rfree: 0.21/0.25

First author:
K. Liu
Resolution: 1.90 Å
R/Rfree: 0.22/0.25
Resolution: 1.90 Å
R/Rfree: 0.22/0.25


First author:
K. Liu
Resolution: 1.95 Å
R/Rfree: 0.22/0.26
Resolution: 1.95 Å
R/Rfree: 0.22/0.26


First author:
M. Lei
Resolution: 2.30 Å
R/Rfree: 0.23/0.27
Resolution: 2.30 Å
R/Rfree: 0.23/0.27


First author:
M. Lei
Resolution: 2.30 Å
R/Rfree: 0.23/0.27
Resolution: 2.30 Å
R/Rfree: 0.23/0.27


First author:
M. Lei
Resolution: 1.84 Å
R/Rfree: 0.21/0.25
Resolution: 1.84 Å
R/Rfree: 0.21/0.25


First author:
K. Liu
Resolution: 2.05 Å
R/Rfree: 0.23/0.27
Resolution: 2.05 Å
R/Rfree: 0.23/0.27

First author:
K. Liu
Resolution: 2.65 Å
R/Rfree: 0.21/0.24
Resolution: 2.65 Å
R/Rfree: 0.21/0.24
X-ray diffraction data for the MBD2 in complex with methylated DNA

First author:
K. Liu
Resolution: 2.15 Å
R/Rfree: 0.21/0.23
Resolution: 2.15 Å
R/Rfree: 0.21/0.23
X-ray diffraction data for the Crystal structure of MBD2 complex with methylated CpG island

First author:
C. Bian
Resolution: 2.10 Å
R/Rfree: 0.20/0.22
Resolution: 2.10 Å
R/Rfree: 0.20/0.22

First author:
C. Xu
Resolution: 1.80 Å
R/Rfree: 0.23/0.25
Resolution: 1.80 Å
R/Rfree: 0.23/0.25
X-ray diffraction data for the Complex of MBD1-MBD and methylated DNA

First author:
K. Liu
Resolution: 2.25 Å
R/Rfree: 0.24/0.27
Resolution: 2.25 Å
R/Rfree: 0.24/0.27
X-ray diffraction data for the Crystal structure of human monoamine oxidase B (MAO B) in complex with fluorophenyl-chromone-carboxamide


First author:
J. Reis
Resolution: 1.70 Å
R/Rfree: 0.16/0.19
Resolution: 1.70 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of human monoamine oxidase B (MAO B) in complex with dimethylphenyl-chromone-carboxamide


First author:
J. Reis
Resolution: 1.80 Å
R/Rfree: 0.17/0.20
Resolution: 1.80 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of a DJ-1 (PARK7) from Homo sapiens at 1.23 A resolution


First author:
Partnership for Nuclear Receptor Signaling Code Biology (NHRs) Joint Center for Structural Genomics (JCSG)
Resolution: 1.23 Å
R/Rfree: 0.12/0.13
Resolution: 1.23 Å
R/Rfree: 0.12/0.13
X-ray diffraction data for the Crystal structure of a ribonucleotide reductase M2 B (RNRR2) from Homo sapiens at 2.20 A resolution


First author:
Partnership for T-Cell Biology (TCELL) Joint Center for Structural Genomics (JCSG)
Resolution: 2.20 Å
R/Rfree: 0.19/0.22
Resolution: 2.20 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of a D-ribulose-5-phosphate-3-epimerase (NP_954699) from HOMO SAPIENS at 2.20 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.20 Å
R/Rfree: 0.16/0.21
Resolution: 2.20 Å
R/Rfree: 0.16/0.21
X-ray diffraction data for the Crystal structure of Putative aminotransferase (AAH25799.1) from MUS MUSCULUS at 1.80 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.80 Å
R/Rfree: 0.17/0.21
Resolution: 1.80 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of Putative aminotransferase (AAH25799.1) from MUS MUSCULUS at 1.65 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.65 Å
R/Rfree: 0.17/0.20
Resolution: 1.65 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of Putative reductase (NP_038806.2) from MUS MUSCULUS at 1.18 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.18 Å
R/Rfree: 0.13/0.16
Resolution: 1.18 Å
R/Rfree: 0.13/0.16
X-ray diffraction data for the Crystal structure of the LnmZ cytochrome P450 hydroxylase from the leinamycin biosynthetic pathway of Streptomyces atroolivaceus S-140 at 1.8 A resolution


First author:
M. Ma
Resolution: 1.80 Å
R/Rfree: 0.16/0.20
Resolution: 1.80 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the Crystal structure of the LnmA cytochrome P450 hydroxylase from the leinamycin biosynthetic pathway of Streptomyces atroolivaceus S-140 at 1.9 A resolution


First author:
M. Ma
Resolution: 1.90 Å
R/Rfree: 0.19/0.22
Resolution: 1.90 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of Deoxyuridine 5'-triphosphate nucleotidohydrolase from Legionella pneumophila Philadelphia 1 in complex with dUMP (Deoxyuridine 5'-monophosphate)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.95 Å
R/Rfree: 0.15/0.18
Resolution: 1.95 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of Deoxyuridine 5'-triphosphate nucleotidohydrolase from Legionella pneumophila Philadelphia 1


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.80 Å
R/Rfree: 0.15/0.18
Resolution: 1.80 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal Structure of Ribose-5-phosphate Isomerase from Legionella pneumophila


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.85 Å
R/Rfree: 0.16/0.22
Resolution: 1.85 Å
R/Rfree: 0.16/0.22
X-ray diffraction data for the Crystal Structure of Ribose-5-phosphate Isomerase from Legionella pneumophila with Bound Substrate Ribose-5-Phosphate and Product Ribulose-5-Phosphate


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 2.00 Å
R/Rfree: 0.15/0.20
Resolution: 2.00 Å
R/Rfree: 0.15/0.20
X-ray diffraction data for the Crystal structure of a 2-dehydro-3-deoxyphosphooctonate aldolase from Legionella pneumophila Philadelphia 1


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 2.55 Å
R/Rfree: 0.18/0.23
Resolution: 2.55 Å
R/Rfree: 0.18/0.23
X-ray diffraction data for the Crystal structure of an organic hydroperoxide resistance protein OsmC, predicted redox protein, regulator of sulfide bond formation from Legionella pneumophila


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Resolution: 1.75 Å
R/Rfree: 0.18/0.21
Resolution: 1.75 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of LdBPK_091320 with inhibitor bound


First author:
Y.H. Lin
Resolution: 2.26 Å
R/Rfree: 0.20/0.24
Resolution: 2.26 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the Crystal structure of L3MBTL1 MBT Domain with MBK14970


First author:
E. DOBROVETSKY
Resolution: 1.74 Å
R/Rfree: 0.20/0.22
Resolution: 1.74 Å
R/Rfree: 0.20/0.22
X-ray diffraction data for the Crystal structure of an EF-Hand calcium binding domain of CAP-Binding Protein Complex-Interacting Protein 1 (EFCAB6) from Homo sapiens at 2.00 A resolution


First author:
Partnership for Nuclear Receptor Signaling Code Biology JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.17/0.20
Resolution: 2.00 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of a nuclear receptor binding factor 2 MIT domain (NRBF2) from Homo sapiens at 1.50 A resolution


First author:
Partnership for Nuclear Receptor Signaling Code Biology (NHRs) JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.50 Å
R/Rfree: 0.21/0.25
Resolution: 1.50 Å
R/Rfree: 0.21/0.25
X-ray diffraction data for the Crystal structure of the KIFC3 motor domain in complex with ADP

First author:
H.W. Park
Resolution: 1.85 Å
R/Rfree: 0.19/0.20
Resolution: 1.85 Å
R/Rfree: 0.19/0.20