6617 results
X-ray diffraction data for the Crystal structure of trypsin at 300 Kelvin with benzamidine

X-ray diffraction data for the Crystal structure of trypsin at 300 Kelvin with benzamidine (duplicate)

X-ray diffraction data for the Crystal structure of the SpyTag003-SpyCatcher003 complex


X-ray diffraction data for the Crystal structure of the allosteric MKP5 mutant Y435W


X-ray diffraction data for the Crystal structure of an MKP5 mutant, Y435W, in complex with an allosteric inhibitor


X-ray diffraction data for the Crystal structure of HP1alpha chromoshadow domain in complex with KAP1 peptide


X-ray diffraction data for the Crystal structure of HP1alpha chromoshadow domain


X-ray diffraction data for the Helicobacter pylori cysteine rich protein B (hcpB)

X-ray diffraction data for the Crystallizing extracellular protein reveals paths for silver mineralization and recovery
First author:
Fiaz Ahmad
X-ray diffraction data for the A Stable Heterotrimeric Foldon with Minimum Mutation


X-ray diffraction data for the Crystal structure of HIV-1 Reverse Transcriptase RNase H domain complexed with a galloyl inhibitor

X-ray diffraction data for the Crystal structure of trypsin at 100 Kelvin with benzamidine (triplicate)

X-ray diffraction data for the Crystal structure of ENL YEATS in complex with histone H3 methacrylated at K18

X-ray diffraction data for the Crystal structure of the H3 hemagglutinin COBRA TJ2


X-ray diffraction data for the Crystal structure of the H3 hemagglutinin COBRA J4
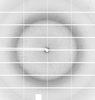


X-ray diffraction data for the Crystal structure of nucleoside-diphosphate kinase from Cryptosporidium parvum in complex with citrate


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: Q5CR64
Resolution: 1.45 Å
R/Rfree: 0.13/0.17
Uniprot: Q5CR64
Resolution: 1.45 Å
R/Rfree: 0.13/0.17
X-ray diffraction data for the Anti-HIV-1 chimeric miniprotein mimicking the N-terminal half of gp41 NHR with an extended region targeting the MPER
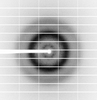


X-ray diffraction data for the The ubiquitin-associated domain of human thirty-eight negative kinase 1, fused to the 3TEL crystallization chaperone via a 2-glycine linker
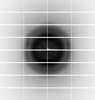


X-ray diffraction data for the Crystal structure of 2,3,4,5-tetrahydropyridine-2,6-dicarboxylate N-succinyltransferase (DapD) from Bordetella pertussis (succinyl-CoA bound)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: P0A4U8
Resolution: 1.45 Å
R/Rfree: 0.16/0.18
Uniprot: P0A4U8
Resolution: 1.45 Å
R/Rfree: 0.16/0.18
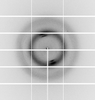
X-ray diffraction data for the Crystal structure of UTP--glucose-1-phosphate uridylyltransferase from Bordetella pertussis


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: Q7VTV0
Resolution: 1.93 Å
R/Rfree: 0.18/0.21
Uniprot: Q7VTV0
Resolution: 1.93 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of nucleoside-diphosphate kinase Cryptosporidium parvum


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: Q5CR64
Resolution: 1.31 Å
R/Rfree: 0.15/0.19
Uniprot: Q5CR64
Resolution: 1.31 Å
R/Rfree: 0.15/0.19
X-ray diffraction data for the Crystal structure of nucleoside-diphosphate kinase from Cryptosporidium parvum in complex with thymidine-5'-phosphate


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: Q5CR64
Resolution: 1.51 Å
R/Rfree: 0.14/0.16
Uniprot: Q5CR64
Resolution: 1.51 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the Crystal structure of nucleoside-diphosphate kinase from Cryptosporidium parvum in complex with cytidine-5'-diphosphate and cytidine-5'-triphosphate


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: Q5CR64
Resolution: 1.61 Å
R/Rfree: 0.14/0.17
Uniprot: Q5CR64
Resolution: 1.61 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of nucleoside-diphosphate kinase from Cryptosporidium parvum in complex with with guanosine-5'-diphosphate and AMP-PNP


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: Q5CR64
Resolution: 1.63 Å
R/Rfree: 0.15/0.17
Uniprot: Q5CR64
Resolution: 1.63 Å
R/Rfree: 0.15/0.17
X-ray diffraction data for the Crystal structure of nucleoside-diphosphate kinase from Cryptosporidium parvum in complex with with ADP and CTP


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: Q5CR64
Resolution: 1.60 Å
R/Rfree: 0.15/0.18
Uniprot: Q5CR64
Resolution: 1.60 Å
R/Rfree: 0.15/0.18