6862 results
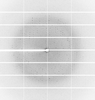
X-ray diffraction data for the Crystal Structure of Tryptophanyl-tRNA synthetase from Klebsiella aerogenes (tryptophan bound)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A0A0H3FKK2
Resolution: 2.15 Å
R/Rfree: 0.18/0.22
Uniprot: A0A0H3FKK2
Resolution: 2.15 Å
R/Rfree: 0.18/0.22
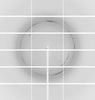
X-ray diffraction data for the Crystal structure of 2,3,4,5-tetrahydropyridine-2,6-dicarboxylate N-succinyltransferase (DapD) from Bordetella pertussis


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: P0A4U8
Resolution: 1.59 Å
R/Rfree: 0.16/0.18
Uniprot: P0A4U8
Resolution: 1.59 Å
R/Rfree: 0.16/0.18
X-ray diffraction data for the Crystal structure of a selenomethionine-labeled hydropyrene synthase (M75L variant) in its closed conformation
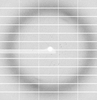


X-ray diffraction data for the Crystal Structure of Tryptophanyl-tRNA synthetase from Klebsiella aerogenes


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A0A0H3FKK2
Resolution: 2.10 Å
R/Rfree: 0.20/0.25
Uniprot: A0A0H3FKK2
Resolution: 2.10 Å
R/Rfree: 0.20/0.25
X-ray diffraction data for the Crystal structure of Phosphoglycerate mutase from Trichomonas vaginalis (sulfate bound)

First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A2DUN8
Resolution: 2.05 Å
R/Rfree: 0.21/0.24
Uniprot: A2DUN8
Resolution: 2.05 Å
R/Rfree: 0.21/0.24
X-ray diffraction data for the Crystal structure of a hydropyrene synthase (M75L variant) in its closed conformation
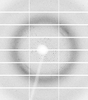


X-ray diffraction data for the Crystal structure of Thymidylate kinase (Tmk) from Klebsiella aerogenes.


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A0A0J2I384
Resolution: 2.85 Å
R/Rfree: 0.24/0.27
Uniprot: A0A0J2I384
Resolution: 2.85 Å
R/Rfree: 0.24/0.27
X-ray diffraction data for the Crystal structure of the Plastid Redox Insensitive 2 from Arabidopsis thaliana
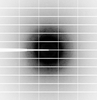


X-ray diffraction data for the Crystal structure of phosphoglycerate mutase from Trichomonas vaginalis
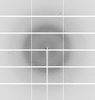


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A2DUN8
Resolution: 1.95 Å
R/Rfree: 0.16/0.19
Uniprot: A2DUN8
Resolution: 1.95 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of bifunctional farnesyl/geranylgeranyl pyrophosphate synthase (FPPS/GGPPS) from Plasmodium falciparum in complex with MMV019313


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: Q8II79
Resolution: 3.25 Å
R/Rfree: 0.25/0.29
Uniprot: Q8II79
Resolution: 3.25 Å
R/Rfree: 0.25/0.29
X-ray diffraction data for the Crystal structure of a mutant Staphylococcus equorum manganese superoxide dismutase T89C and N187C
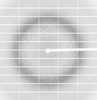


X-ray diffraction data for the Crystal structure of AF9 YEATS domain in complex with acetylated at K1007 MOZ


First author:
K. Selvam
Resolution: 2.30 Å
R/Rfree: 0.25/0.27
Resolution: 2.30 Å
R/Rfree: 0.25/0.27
X-ray diffraction data for the Crystal structure of AF9 YEATS domain in complex with dicrotonylated at K1007 and K1014 MOZ


First author:
K. Selvam
Resolution: 2.10 Å
R/Rfree: 0.20/0.25
Resolution: 2.10 Å
R/Rfree: 0.20/0.25
X-ray diffraction data for the Crystal structure of Purine nucleoside phosphorylase from Trichomonas vaginalis (inosine bound)

First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Resolution: 1.20 Å
R/Rfree: 0.13/0.16
Resolution: 1.20 Å
R/Rfree: 0.13/0.16
X-ray diffraction data for the Crystal structure of a bacterial clusterless MutYX bound to an Abasic site analog (THF) opposite d(8-oxo-G)


First author:
C.H. Trasvina-Arenas
Resolution: 1.55 Å
R/Rfree: 0.18/0.20
Resolution: 1.55 Å
R/Rfree: 0.18/0.20
X-ray diffraction data for the Heligmosomoides polygyrus TGF-beta Mimic 6 Domain 3 (TGM6-D3) Bound to Human TGF-beta Type II Receptor Extracellular Domain
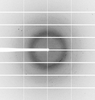


First author:
S.E. White
Resolution: 1.40 Å
R/Rfree: 0.16/0.18
Resolution: 1.40 Å
R/Rfree: 0.16/0.18
X-ray diffraction data for the Crystal structure of HIV-1 capsid N-terminal domain in the presence of Lenacapavir
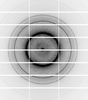


X-ray diffraction data for the Crystal structure of Purine nucleoside phosphorylase from Trichomonas vaginalis (adenosine and phosphate bound)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A2EU62
Resolution: 1.24 Å
R/Rfree: 0.13/0.15
Uniprot: A2EU62
Resolution: 1.24 Å
R/Rfree: 0.13/0.15
X-ray diffraction data for the Crystal structure of human leukocyte antigen A*0101 in complex with the Fab of alloreactive antibody E07


X-ray diffraction data for the Crystal structure of AtNATA1 bound to Ornithine and CoA


X-ray diffraction data for the Human astrovirus 2 spike bound to neutralizing scFv 4B6
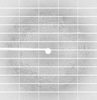

First author:
S. Lanning
Resolution: 2.67 Å
R/Rfree: 0.21/0.26
Resolution: 2.67 Å
R/Rfree: 0.21/0.26
X-ray diffraction data for the Crystal structure of Purine nucleoside phosphorylase from Trichomonas vaginalis (C2 form)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A2EU62
Resolution: 1.80 Å
R/Rfree: 0.16/0.20
Uniprot: A2EU62
Resolution: 1.80 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the Crystal structure of Purine nucleoside phosphorylase from Trichomonas vaginalis (P21 form)
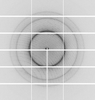


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A2EU62
Resolution: 1.59 Å
R/Rfree: 0.15/0.18
Uniprot: A2EU62
Resolution: 1.59 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of Purine nucleoside phosphorylase from Trichomonas vaginalis (Hepes bound)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID) Seattle Structural Genomics Center for Infectious Disease
Uniprot: A2EU62
Resolution: 1.49 Å
R/Rfree: 0.15/0.17
Uniprot: A2EU62
Resolution: 1.49 Å
R/Rfree: 0.15/0.17
X-ray diffraction data for the Tetramer Formation of the BCL11A ZF0 Domain

