1390 results
X-ray diffraction data for the Crystal structure of 3-ketoacyl-(acyl-carrier-protein) reductase from Brucella melitensis


First author:
T.E. Edwards
Resolution: 2.20 Å
R/Rfree: 0.17/0.23
Resolution: 2.20 Å
R/Rfree: 0.17/0.23
X-ray diffraction data for the Crystal structure of ribose-5-phosphate isomerase B RpiB from Giardia lamblia


First author:
T.E. Edwards
Resolution: 2.30 Å
R/Rfree: 0.20/0.23
Resolution: 2.30 Å
R/Rfree: 0.20/0.23
X-ray diffraction data for the Crystal structure of a dephospho-CoA kinase from Burkholderia vietnamiensis bound to ADP


First author:
T.E. Edwards
Resolution: 2.60 Å
R/Rfree: 0.21/0.25
Resolution: 2.60 Å
R/Rfree: 0.21/0.25
X-ray diffraction data for the Structure of a putative esterase Rv1847/MT1895 from Mycobacterium tuberculosis


First author:
M.C. Clifton
Resolution: 1.70 Å
R/Rfree: 0.17/0.20
Resolution: 1.70 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of Adenylosuccinate synthetase from Legionella pneumophila Philadelphia 1


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.95 Å
R/Rfree: 0.19/0.23
Resolution: 1.95 Å
R/Rfree: 0.19/0.23
X-ray diffraction data for the Crystal structure of hemagglutinin from H1N1 Influenza A virus A/Denver/57 bound to the C05 antibody


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 2.92 Å
R/Rfree: 0.20/0.26
Resolution: 2.92 Å
R/Rfree: 0.20/0.26
X-ray diffraction data for the Crystal structure of a malonate decarboxylase, alpha subunit from Acinetobacter baumannii


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.70 Å
R/Rfree: 0.16/0.20
Resolution: 1.70 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the Crystal structure of InvbP.18715.a.KN11: Influenza hemagglutinin from strain A/Almaty/32/1998


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 2.05 Å
R/Rfree: 0.19/0.22
Resolution: 2.05 Å
R/Rfree: 0.19/0.22
X-ray diffraction data for the Crystal structure of an dephospho-CoA kinase CoaE from Mycobacterium paratuberculosis


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 2.15 Å
R/Rfree: 0.17/0.19
Resolution: 2.15 Å
R/Rfree: 0.17/0.19
X-ray diffraction data for the Crystal structure of an alcohol dehydrogenase from Elizabethkingia anophelis NUHP1


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 2.10 Å
R/Rfree: 0.15/0.19
Resolution: 2.10 Å
R/Rfree: 0.15/0.19
X-ray diffraction data for the Crystal structure of fumarate reductase, flavo protein subunit, from Helicobacter pylori G27


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 2.35 Å
R/Rfree: 0.17/0.22
Resolution: 2.35 Å
R/Rfree: 0.17/0.22
X-ray diffraction data for the CRYSTAL STRUCTURE OF ENOLASE FROM LEGIONELLA PNEUMOPHILA BOUND TO PHOSPHATE AND MAGNESIUM


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.90 Å
R/Rfree: 0.16/0.20
Resolution: 1.90 Å
R/Rfree: 0.16/0.20
X-ray diffraction data for the Crystal structure of 1,4-dihydroxy-2-naphthoyl-CoA synthase Elizabethkingia anophelis NUHP1


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.60 Å
R/Rfree: 0.15/0.18
Resolution: 1.60 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Structure of DNA polymerase III subunit beta from Borrelia burgdorferi in complex with a natural product


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 2.05 Å
R/Rfree: 0.20/0.24
Resolution: 2.05 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the Crystal Structure of CiaD from Campylobacter jejuni (C-terminal fragment)

First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 2.45 Å
R/Rfree: 0.22/0.27
Resolution: 2.45 Å
R/Rfree: 0.22/0.27
X-ray diffraction data for the Crystal structure of alpha-glucosidase (yicI) from Klebsiella aerogenes

First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Uniprot: A0A0H3FLJ3
Resolution: 2.70 Å
R/Rfree: 0.23/0.26
Uniprot: A0A0H3FLJ3
Resolution: 2.70 Å
R/Rfree: 0.23/0.26
X-ray diffraction data for the Crystal Structure of Dephospho-CoA kinase from Klebsiella aerogenes (ADP Bound)

First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Uniprot: A0A0H3FR62
Resolution: 1.40 Å
R/Rfree: 0.16/0.19
Uniprot: A0A0H3FR62
Resolution: 1.40 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal Structure of Dephospho-CoA kinase from Klebsiella aerogenes (ATP bound)
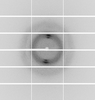


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Uniprot: A0A0H3FR62
Resolution: 1.50 Å
R/Rfree: 0.17/0.20
Uniprot: A0A0H3FR62
Resolution: 1.50 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of N-acetylneuraminate lyase (NanA) from Klebsiella aerogenes (pyruvate bound, Orthorhombic P form)

First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 2.10 Å
R/Rfree: 0.15/0.19
Resolution: 2.10 Å
R/Rfree: 0.15/0.19
X-ray diffraction data for the Crystal structure of MurC from Pseudomonas aeruginosa in complex (sulfate bound)

First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Uniprot: Q9HW02
Resolution: 2.32 Å
R/Rfree: 0.20/0.21
Uniprot: Q9HW02
Resolution: 2.32 Å
R/Rfree: 0.20/0.21
X-ray diffraction data for the Crystal structure of glutamate dehydrogenase from Babesia microti


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Uniprot: A0A0K3AUK4
Resolution: 2.18 Å
R/Rfree: 0.18/0.21
Uniprot: A0A0K3AUK4
Resolution: 2.18 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of a Heat shock protein (DnaJ) from Brucella melitensis


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Uniprot: Q2YMC9
Resolution: 3.09 Å
R/Rfree: 0.24/0.29
Uniprot: Q2YMC9
Resolution: 3.09 Å
R/Rfree: 0.24/0.29
X-ray diffraction data for the Structure of a Glycosyltransferase from Leptospira borgpetersenii serovar Hardjo-bovis (strain JB197)
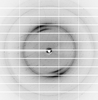


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.95 Å
R/Rfree: 0.15/0.18
Resolution: 1.95 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal Structure of Enoyl-[acyl-carrier-protein] reductase from Mycobacterium avium with bound NAD


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.65 Å
R/Rfree: 0.14/0.17
Resolution: 1.65 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of a Probable carnitine operon oxidoreductase caia from Brucella melitensis


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 2.60 Å
R/Rfree: 0.20/0.24
Resolution: 2.60 Å
R/Rfree: 0.20/0.24