1469 results
X-ray diffraction data for the Crystal structure of a hepn domain containing protein (af_0298) from archaeoglobus fulgidus at 1.95 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.95 Å
R/Rfree: 0.18/0.21
Resolution: 1.95 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of an EF-Hand calcium binding domain of CAP-Binding Protein Complex-Interacting Protein 1 (EFCAB6) from Homo sapiens at 2.00 A resolution


First author:
Partnership for Nuclear Receptor Signaling Code Biology JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.17/0.20
Resolution: 2.00 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of a SusD-like protein (BT_3752) from Bacteroides thetaiotaomicron VPI-5482 at 1.70 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.70 Å
R/Rfree: 0.14/0.17
Resolution: 1.70 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of a putative hydrolase (ava_4197) from anabaena variabilis atcc 29413 at 1.35 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.35 Å
R/Rfree: 0.12/0.14
Resolution: 1.35 Å
R/Rfree: 0.12/0.14
X-ray diffraction data for the Crystal structure of an ntf2-like protein with a cystatin-like fold (saro_3722) from novosphingobium aromaticivorans dsm at 1.16 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.16 Å
R/Rfree: 0.14/0.17
Resolution: 1.16 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of an rmlc-like cupin protein (reut_a0381) from ralstonia eutropha jmp134 at 2.60 A resolution

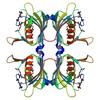
First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.60 Å
R/Rfree: 0.23/0.26
Resolution: 2.60 Å
R/Rfree: 0.23/0.26
X-ray diffraction data for the Crystal structure of Pyridoxamine 5'-phosphate oxidase-like protein (NP_783940.1) from Lactobacillus plantarum at 1.55 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.55 Å
R/Rfree: 0.17/0.20
Resolution: 1.55 Å
R/Rfree: 0.17/0.20
X-ray diffraction data for the Crystal structure of a DUF2790 family protein (PA3229) from Pseudomonas aeruginosa PAO1 at 1.26 A resolution

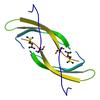
First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.26 Å
R/Rfree: 0.14/0.17
Resolution: 1.26 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal structure of Dihydrodipicolinate synthase (TM1521) from Thermotoga maritima at 1.80 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.80 Å
R/Rfree: 0.14/0.19
Resolution: 1.80 Å
R/Rfree: 0.14/0.19
X-ray diffraction data for the CRYSTAL STRUCTURE OF A RMLC-LIKE CUPIN (SFRI_3105) FROM SHEWANELLA FRIGIDIMARINA NCIMB 400 AT 1.90 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.19/0.26
Resolution: 1.90 Å
R/Rfree: 0.19/0.26
X-ray diffraction data for the Crystal structure of a putative AphA-like transcription factor (ZP_00208345.1) from Magnetospirillum magnetotacticum MS-1 at 2.00 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.21/0.26
Resolution: 2.00 Å
R/Rfree: 0.21/0.26
X-ray diffraction data for the Crystal structure of a Putative aldose 1-epimerase (b2544) from ESCHERICHIA COLI K12 at 1.59 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.59 Å
R/Rfree: 0.16/0.19
Resolution: 1.59 Å
R/Rfree: 0.16/0.19
X-ray diffraction data for the Crystal structure of a DUF4468 family protein (BACOVA_04320) from Bacteroides ovatus ATCC 8483 at 1.49 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.49 Å
R/Rfree: 0.16/0.18
Resolution: 1.49 Å
R/Rfree: 0.16/0.18
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE HYDROLASE (BVU_4111) FROM BACTEROIDES VULGATUS ATCC 8482 AT 1.90 A RESOLUTION


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.17/0.21
Resolution: 1.90 Å
R/Rfree: 0.17/0.21
X-ray diffraction data for the Crystal structure of a putative GDSL-like lipase (BACUNI_00748) from Bacteroides uniformis ATCC 8492 at 1.90 A resolution


First author:
JOINT CENTER FOR STRUCTURAL GENOMICS (JCSG)
Resolution: 1.90 Å
R/Rfree: 0.16/0.21
Resolution: 1.90 Å
R/Rfree: 0.16/0.21
X-ray diffraction data for the Crystal structure of a dinb-like protein (bce_4655) from bacillus cereus atcc 10987 at 2.01 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.01 Å
R/Rfree: 0.23/0.27
Resolution: 2.01 Å
R/Rfree: 0.23/0.27
X-ray diffraction data for the Crystal structure of Imidazole glycerol phosphate synthase subunit hisF (EC 4.1.3.-) (tm1036) from Thermotoga maritima at 1.64 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.64 Å
R/Rfree: 0.17/0.19
Resolution: 1.64 Å
R/Rfree: 0.17/0.19
X-ray diffraction data for the Crystal structure of a Putative oxygenase (SO_2589) from Shewanella oneidensis at 2.20 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.20 Å
R/Rfree: 0.19/0.24
Resolution: 2.20 Å
R/Rfree: 0.19/0.24
X-ray diffraction data for the Crystal structure of a SusD homolog (BDI_0600) from Parabacteroides distasonis ATCC 8503 AT 2.05 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.05 Å
R/Rfree: 0.24/0.27
Resolution: 2.05 Å
R/Rfree: 0.24/0.27
X-ray diffraction data for the CRYSTAL STRUCTURE OF A PUTATIVE THIOESTERASE (KT2440) FROM PSEUDOMONAS PUTIDA KT2440 AT 2.00 A RESOLUTION

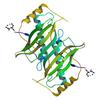
First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.00 Å
R/Rfree: 0.17/0.23
Resolution: 2.00 Å
R/Rfree: 0.17/0.23
X-ray diffraction data for the Crystal structure of a putative adhesin (BF0245) from Bacteroides fragilis NCTC 9343 at 2.07 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.07 Å
R/Rfree: 0.18/0.21
Resolution: 2.07 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of Putative Oxygenase (YP_001051978.1) from SHEWANELLA BALTICA OS155 at 2.26 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.26 Å
R/Rfree: 0.20/0.23
Resolution: 2.26 Å
R/Rfree: 0.20/0.23
X-ray diffraction data for the Crystal structure of a D-ribulose-5-phosphate-3-epimerase (NP_954699) from HOMO SAPIENS at 2.20 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 2.20 Å
R/Rfree: 0.16/0.21
Resolution: 2.20 Å
R/Rfree: 0.16/0.21
X-ray diffraction data for the Crystal structure of a putative thioesterase (tm1040_2492) from silicibacter sp. tm1040 at 1.79 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.79 Å
R/Rfree: 0.22/0.27
Resolution: 1.79 Å
R/Rfree: 0.22/0.27
X-ray diffraction data for the Crystal structure of BT_1490 (NP_810393.1) from BACTEROIDES THETAIOTAOMICRON VPI-5482 at 1.50 A resolution


First author:
Joint Center for Structural Genomics (JCSG)
Resolution: 1.50 Å
R/Rfree: 0.17/0.20
Resolution: 1.50 Å
R/Rfree: 0.17/0.20