1390 results
X-ray diffraction data for the Crystal structure of a betaine aldehyde dehydrogenase from Burkholderia pseudomallei bound to cofactor NAD


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.55 Å
R/Rfree: 0.14/0.17
Resolution: 1.55 Å
R/Rfree: 0.14/0.17
X-ray diffraction data for the Crystal Structure of aspartate-semialdehyde dehydrogenase from Acinetobacter baumannii in complex with NADP


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.95 Å
R/Rfree: 0.15/0.18
Resolution: 1.95 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal Structure of UDP-N-acetylmuramoylalanine-D-glutamate ligase from Acinetobacter baumannii AB5075-UW in complex with ADP


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.50 Å
R/Rfree: 0.15/0.17
Resolution: 1.50 Å
R/Rfree: 0.15/0.17
X-ray diffraction data for the Crystal Structure of Methionine-tRNA ligase / Methionyl-tRNA synthetase (MetRS) from Pseudomonas aeruginosa PAO1


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 2.80 Å
R/Rfree: 0.18/0.20
Resolution: 2.80 Å
R/Rfree: 0.18/0.20
X-ray diffraction data for the Crystal structure of phosphoprotein/Protein P/Protein M1 residues 69-297 from Rabies virus reveals degradation to C-terminal domain only

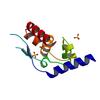
First author:
T.E. Edwards
Resolution: 2.20 Å
R/Rfree: 0.17/0.23
Resolution: 2.20 Å
R/Rfree: 0.17/0.23
X-ray diffraction data for the Crystal structure of nucleoside diphosphate kinase from Giardia lamblia featuring a disordered dinucleotide binding site


First author:
T.E. Edwards
Resolution: 2.65 Å
R/Rfree: 0.23/0.27
Resolution: 2.65 Å
R/Rfree: 0.23/0.27
X-ray diffraction data for the Crystal structure of Glutamate-1-semialdehyde 2,1- aminomutase (GSA) from Pseudomonas aeruginosa


First author:
Delker Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.75 Å
R/Rfree: 0.15/0.18
Resolution: 1.75 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal Structure of Prolyl-tRNA Synthetase from Cryptosporidium parvum complexed with L-Proline and AMP


First author:
Dranow Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 2.80 Å
R/Rfree: 0.21/0.24
Resolution: 2.80 Å
R/Rfree: 0.21/0.24
X-ray diffraction data for the Crystal Structure of Lysyl-tRNA Synthetase from Cryptosporidium parvum complexed with L-lysine and cladosporin


First author:
Dranow Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.90 Å
R/Rfree: 0.20/0.25
Resolution: 1.90 Å
R/Rfree: 0.20/0.25
X-ray diffraction data for the Crystal structure of a Putative aldehyde dehydrogenase family protein Burkholderia cenocepacia J2315 in complex with partially reduced NADH


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 2.10 Å
R/Rfree: 0.15/0.19
Resolution: 2.10 Å
R/Rfree: 0.15/0.19
X-ray diffraction data for the Crystal structure of a glucose-1-phosphate thymidylyltransferase from Burkholderia phymatum bound to 2'-deoxy-thymidine-B-L-rhamnose


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 2.10 Å
R/Rfree: 0.18/0.23
Resolution: 2.10 Å
R/Rfree: 0.18/0.23
X-ray diffraction data for the Crystal Structure of Ribose-phosphate Pyrophosphokinase from Legionella pneumophila with bound AMP, ADP, and Ribose-5-Phosphate


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.75 Å
R/Rfree: 0.15/0.18
Resolution: 1.75 Å
R/Rfree: 0.15/0.18
X-ray diffraction data for the Crystal structure of prolyl-tRNA synthetase from Naegleria fowleri in complex with proline and adenosine monophophsphate (AMP)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 2.00 Å
R/Rfree: 0.15/0.19
Resolution: 2.00 Å
R/Rfree: 0.15/0.19
X-ray diffraction data for the Crystal structure of Bifunctional adenosylcobalamin biosynthesis protein from Klebsiella pneumoniae


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.65 Å
R/Rfree: 0.18/0.21
Resolution: 1.65 Å
R/Rfree: 0.18/0.21
X-ray diffraction data for the Crystal structure of 2,3-diketo-5-methylthiopentyl-1-phosphate enolase-phosphatase from Klebsiella aerogenes (P1 Form)

First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 2.00 Å
R/Rfree: 0.19/0.25
Resolution: 2.00 Å
R/Rfree: 0.19/0.25
X-ray diffraction data for the Crystal Structure of Acetyl-CoA synthetase in complex with an allyl ester AMP inhibitor from Cryptococcus neoformans H99

First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 2.65 Å
R/Rfree: 0.18/0.24
Resolution: 2.65 Å
R/Rfree: 0.18/0.24
X-ray diffraction data for the Crystal Structure of Histidinol-phosphate aminotransferase from Klebsiella pneumoniae subsp. pneumoniae (strain HS11286)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.80 Å
R/Rfree: 0.15/0.19
Resolution: 1.80 Å
R/Rfree: 0.15/0.19
X-ray diffraction data for the Crystal Structure of Cystathionine beta lyase from Klebsiella aerogenes, PLP-Oxamate Adduct (C2 form)

First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.10 Å
R/Rfree: 0.12/0.14
Resolution: 1.10 Å
R/Rfree: 0.12/0.14
X-ray diffraction data for the Crystal Structure of Cystathionine beta lyase from Klebsiella aerogenes, Covalently bound and free PLP (I2 form)
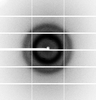

First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.30 Å
R/Rfree: 0.13/0.15
Resolution: 1.30 Å
R/Rfree: 0.13/0.15
X-ray diffraction data for the Crystal Structure of Cystathionine beta lyase from Klebsiella aerogenes, PLP and phosphate bound (C2 form)
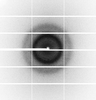

First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.45 Å
R/Rfree: 0.14/0.16
Resolution: 1.45 Å
R/Rfree: 0.14/0.16
X-ray diffraction data for the Crystal Structure of Cystathionine beta lyase from Klebsiella aerogenes, PLP adduct with Alanine (C2 form)
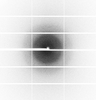

First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.60 Å
R/Rfree: 0.16/0.18
Resolution: 1.60 Å
R/Rfree: 0.16/0.18
X-ray diffraction data for the Crystal Structure of Cystathionine beta lyase from Klebsiella aerogenes, PLP/Malonate complex (C2 form)
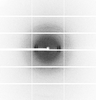

First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.40 Å
R/Rfree: 0.15/0.16
Resolution: 1.40 Å
R/Rfree: 0.15/0.16
X-ray diffraction data for the Crystal Structure of ArnB Transferase from Klebsiella aerogenes (Lattice Translocation Disorder, P21 form)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.55 Å
R/Rfree: 0.20/0.22
Resolution: 1.55 Å
R/Rfree: 0.20/0.22
X-ray diffraction data for the Crystal Structure of ArnB Transferase from Klebsiella aerogenes (Lattice Translocation Disorder, P1 form)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.75 Å
R/Rfree: 0.20/0.24
Resolution: 1.75 Å
R/Rfree: 0.20/0.24
X-ray diffraction data for the Crystal Structure of ArnB Transferase from Klebsiella aerogenes (Lattice Translocation Disorder, P1 form2)


First author:
Seattle Structural Genomics Center for Infectious Disease (SSGCID)
Resolution: 1.50 Å
R/Rfree: 0.15/0.18
Resolution: 1.50 Å
R/Rfree: 0.15/0.18